Analyse de données métagénomiques 16S - FROGS
Master BMC - Atelier Biodiversité microbienne
September 22, 2023
Missions de la plateforme Migale
- Mettre à disposition une infrastructure de calcul scientifique pour la génomique
- Calcul / Stockage / Outils / Données
- Diffuser un savoir-faire en bioinformatique et biostatistique
- Formations / Assistance / Conseil
- Concevoir et développer des applications
- Développement et mise à disposition d’outils / Interfaces innovantes
- Analyser des données génomiques
- Génomique / Génomique comparée / Métagénomique
Une Infrastructure environnée pour la bioinformatique
- Une cinquantaine de serveurs physiques et virtuels hébergés au DataCenter INRAE Île de France
- 819 utilisateurs actifs début 2021 ( 219 nvx comptes)
- Cluster de calcul
- 2660 cœurs , 13,5 To de RAM
- 1 machine highmem 864 Go
- 1 machine bigmem 2 To
- 1,7 M calculs lancés en 2022, ~303 années de calcul ( 205 en 2021)
- Espace de stockage centralisé de 712 To (55% occupation)
Une Infrastructure environnée pour la bioinformatique
- Plus de 500 outils et packages bioinformatiques installés (ligne de commande, Galaxy, R…)
- 80 banques de données publiques mises à disposition, formatées et mises à jour automatiquement
- Différents types d’accès selon les utilisateurs et les besoins :
- SSH (ligne de commande) pour l’accès aux ressource de calcul
- Galaxy (interface web, workflows) pour les utilisateurs non experts
- Rstudio (interface web) pour les analyses stats
- Des interfaces web dédiées à certains types d’analyse
FROGS
What is FROGS

- FROGS is a software package for accurate, simple and robust processing of metabarcoding sequencing reads.
- FROGS uses standard methods and tools combined with original and innovative approaches
- FROGS currently offers 29 tools, numerous graphs, statistics and functional inference, providing biologists with enhanced support for their analyses.
- FROGS use is open to both novices and experts, thanks to the ability to launch tools via the Galaxy platforms or via the command line.
FROGS in brief
- 2 publications
- 20,000+ downloads worldwide
- 350+ scientists have completed our training courses
- 550+ citations
- 80+ databanks available for taxonomic affiliation
FROGS features
- Able to deal with:
- All amplicons (whatever their size)!
- Short and long reads
- Merged and unmerged reads
FROGS team
- FROGS is a INRAE collaborative project since 2015


FROGS articles


How to use FROGS

- Command line
remove_chimera.py
--input-biom clustering.biom \
--input-fasta clustering.fasta \
--non-chimera remove_chimera.fasta \
--out-abundance remove_chimera.biom \
--summary remove_chimera.html
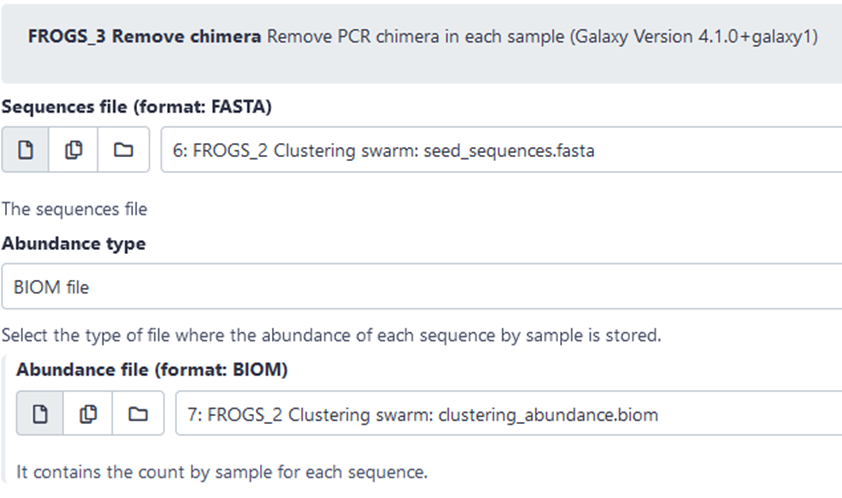
- Galaxy instances via web
![]()
FROGS 16S SOP
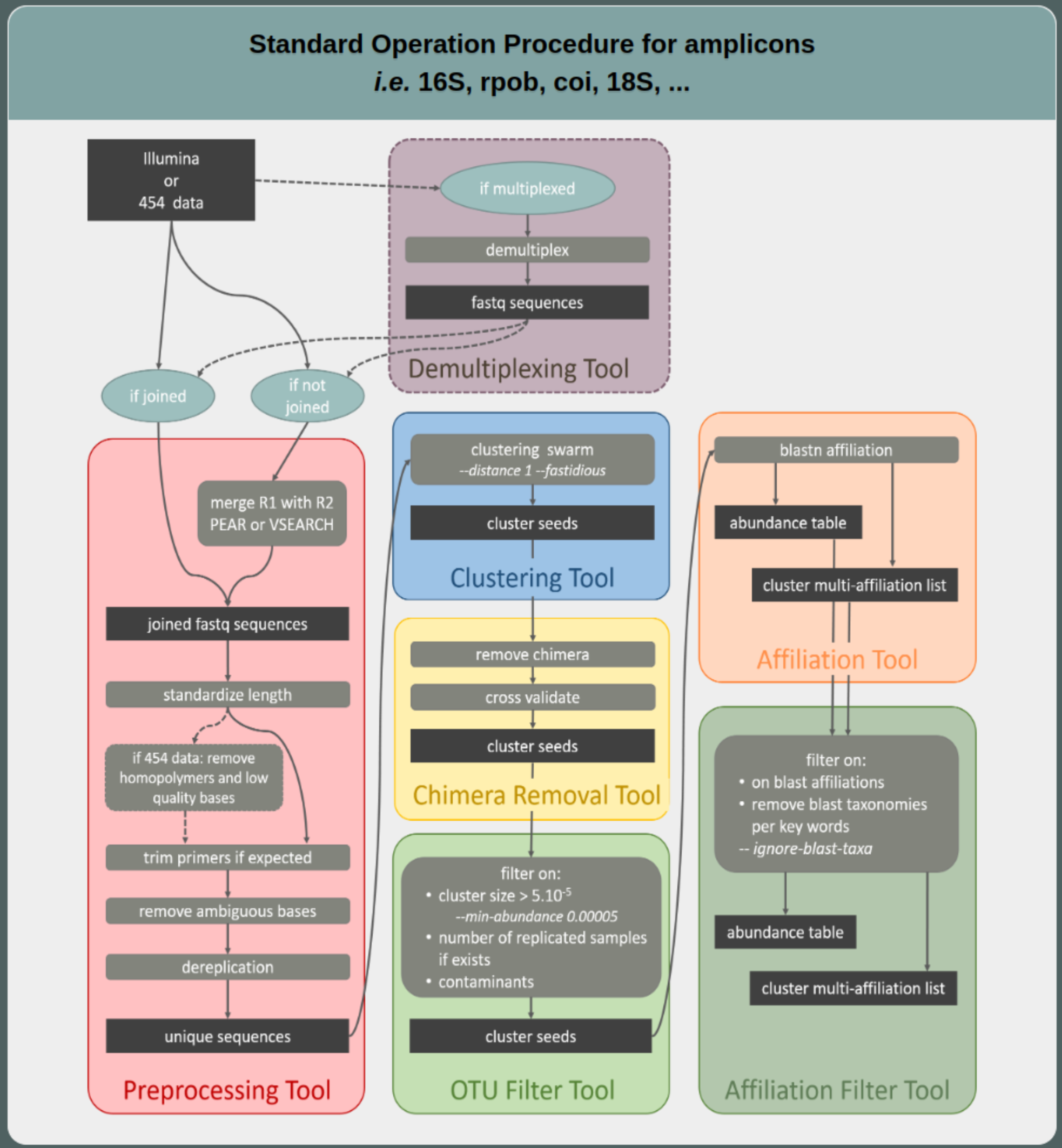
FROGS docs and help

- Website: https://frogs.toulouse.inrae.fr
- Github: https://github.com/geraldinepascal/FROGS.git
- Newsletter: subscription request at frogs-support@inrae.fr
- Need help
- frogs-support@inrae.fr for generic questions
- help-migale@inrae.fr for bugs/quotas/errors with Galaxy Migale instance
Sequencing data
FASTQ format
@ST-E00114:1342:HHMGVCCX2:1:1101:3123:2012 1:N:0:TCCGGAGA+TCAGAGCC
CTTGGTCATTTAGAG
+
***<<*AEF???***
@ST-E00114:1342:HHMGVCCX2:1:1101:11556:2030 1:N:0:TCCGGAGA+TCAGAGCC
CATTGGCCATATCAT
+
AAAE??<<*???***Meaning
@Identifier1 (comment)
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
+
QQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQ
@Identifier2 (comment)
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
+
QQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQuality score encoding
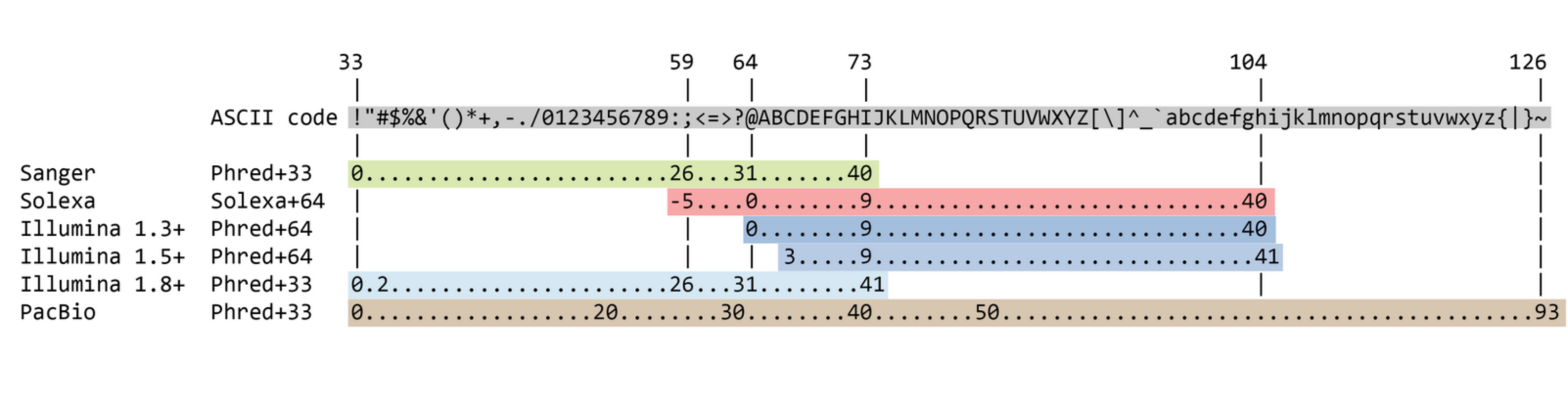
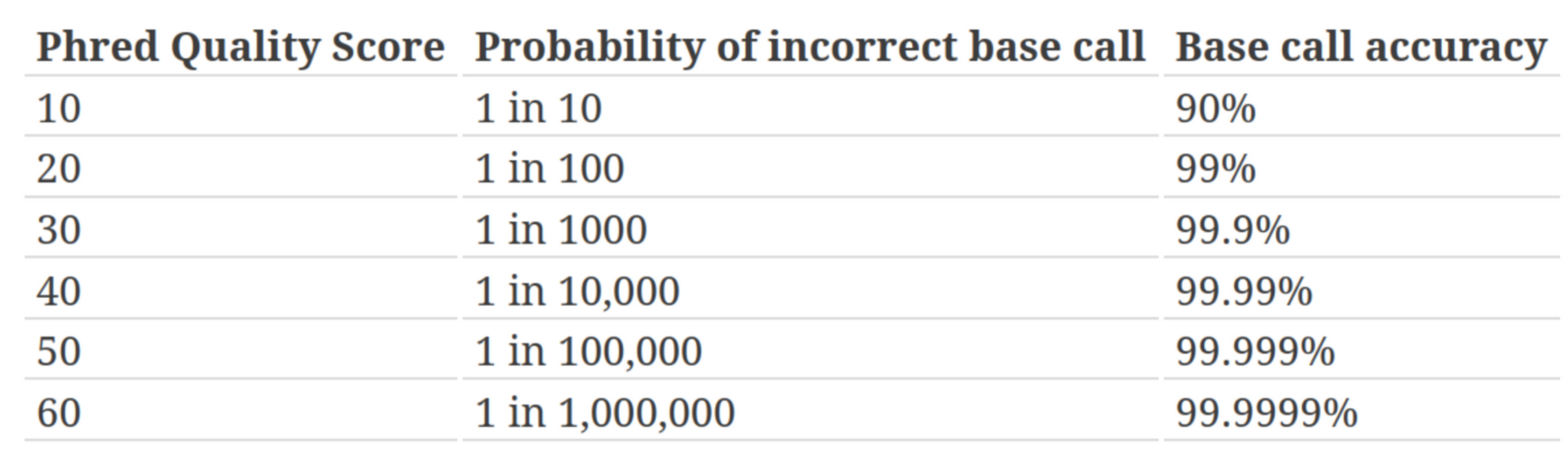
Quality score
Measure of the quality of the identification of the nucleobases generated by automated DNA sequencing
FASTQ compression
- Compression is essential to deal with FASTQ files (reduce disk storage)
- extension:
file.fastq.gz - Tools are (almost all) able to deal with compressed files
Quality control
Quality control
- One of the most easy step in bioinformatics …
- … but one of the most important
- check if everything is ok
- Indicates if/how to clean reads
- Shows possible sequencing problems
- The results must be interpreted in relation to what has been sequenced
Reads are not perfect
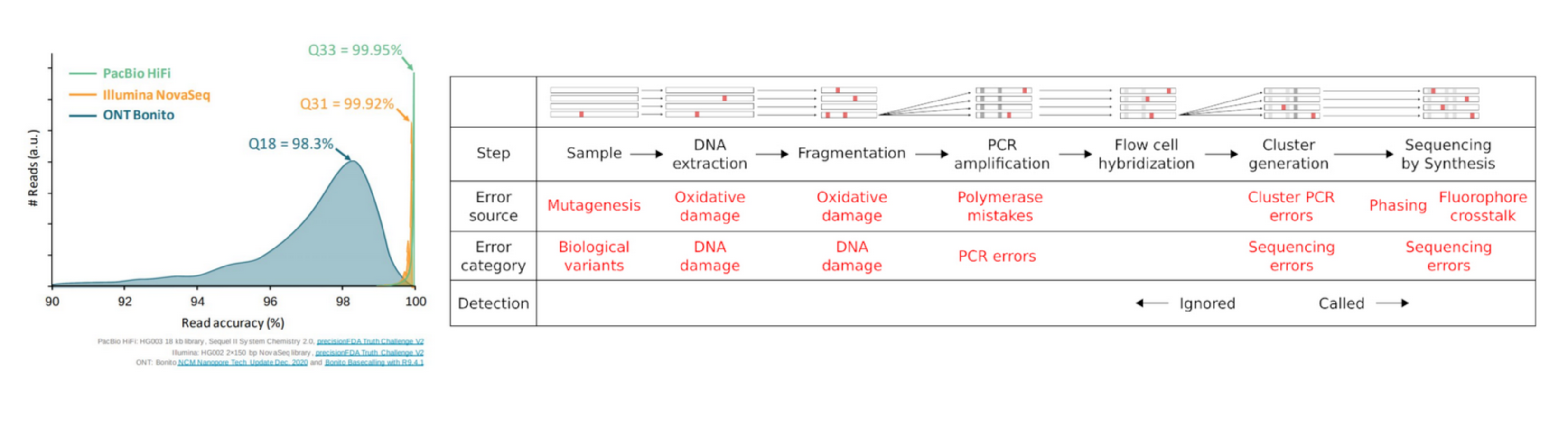
Why QC’ing your reads?
Try to answer to (not always) simple questions:
- Are data conform to the expected level of performance?
- Size / Number of reads / Quality
- Residual presence of adapters or indexes?
- (Un)expected techincal biases?
- (Un)expected biological biases?
Warning
QC without context leads to misinterpretation!
Demultiplexing

Multiplexing principle
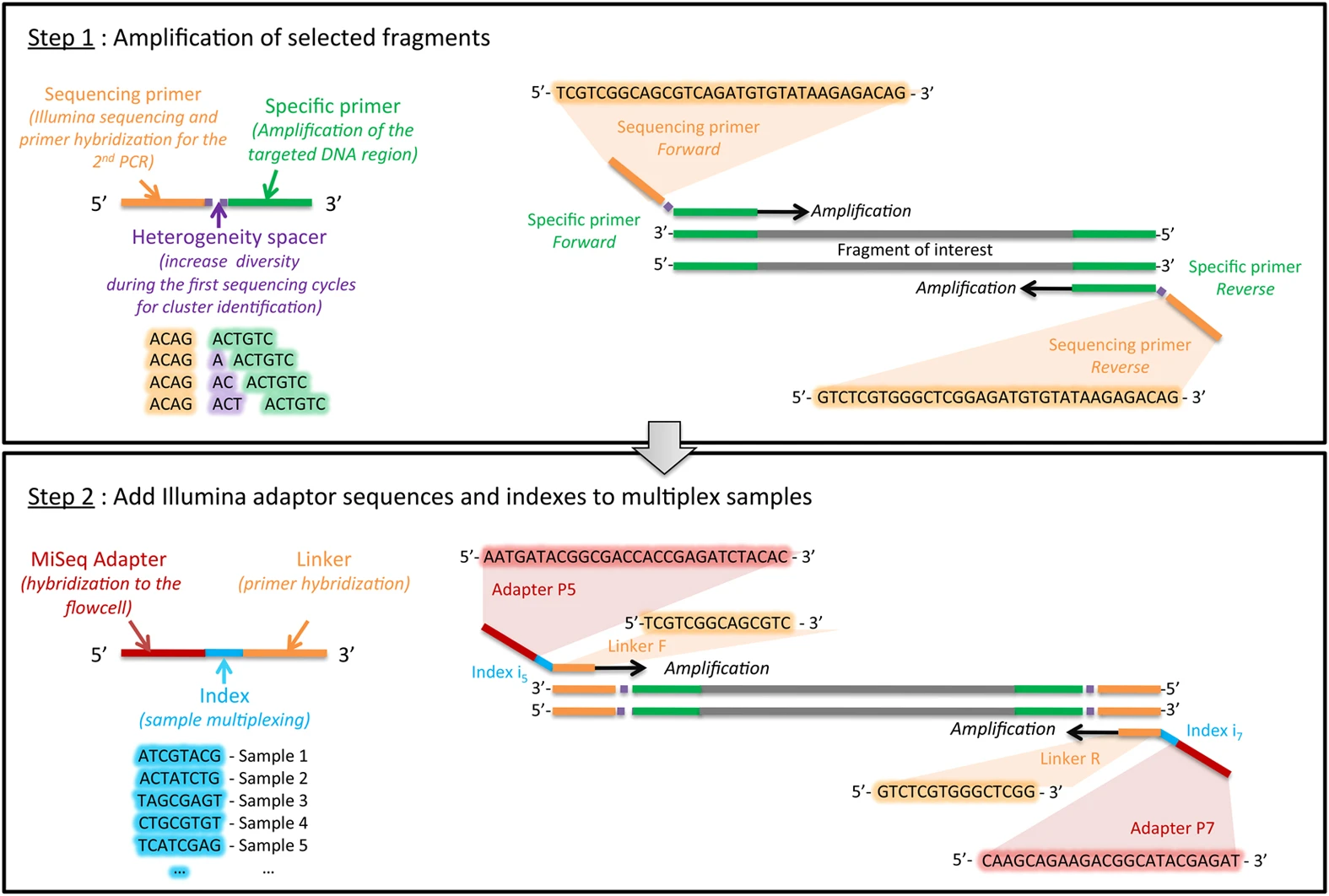
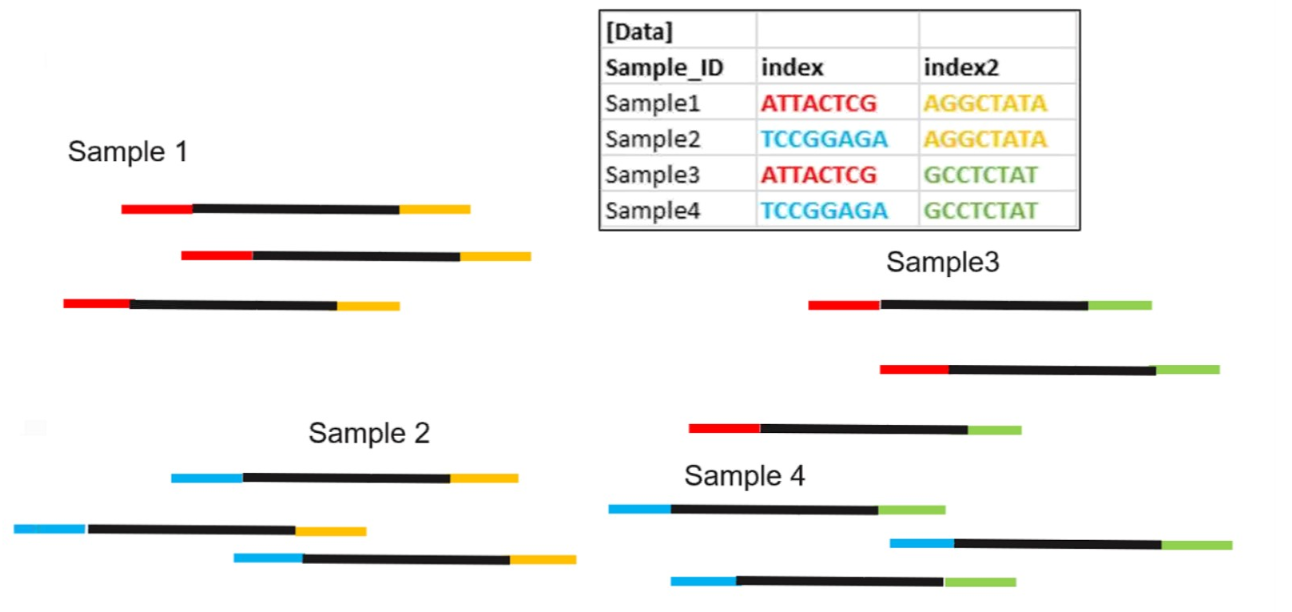
Demultiplexing by bioinformatics
FROGS preprocess

FROGS preprocess deals with…
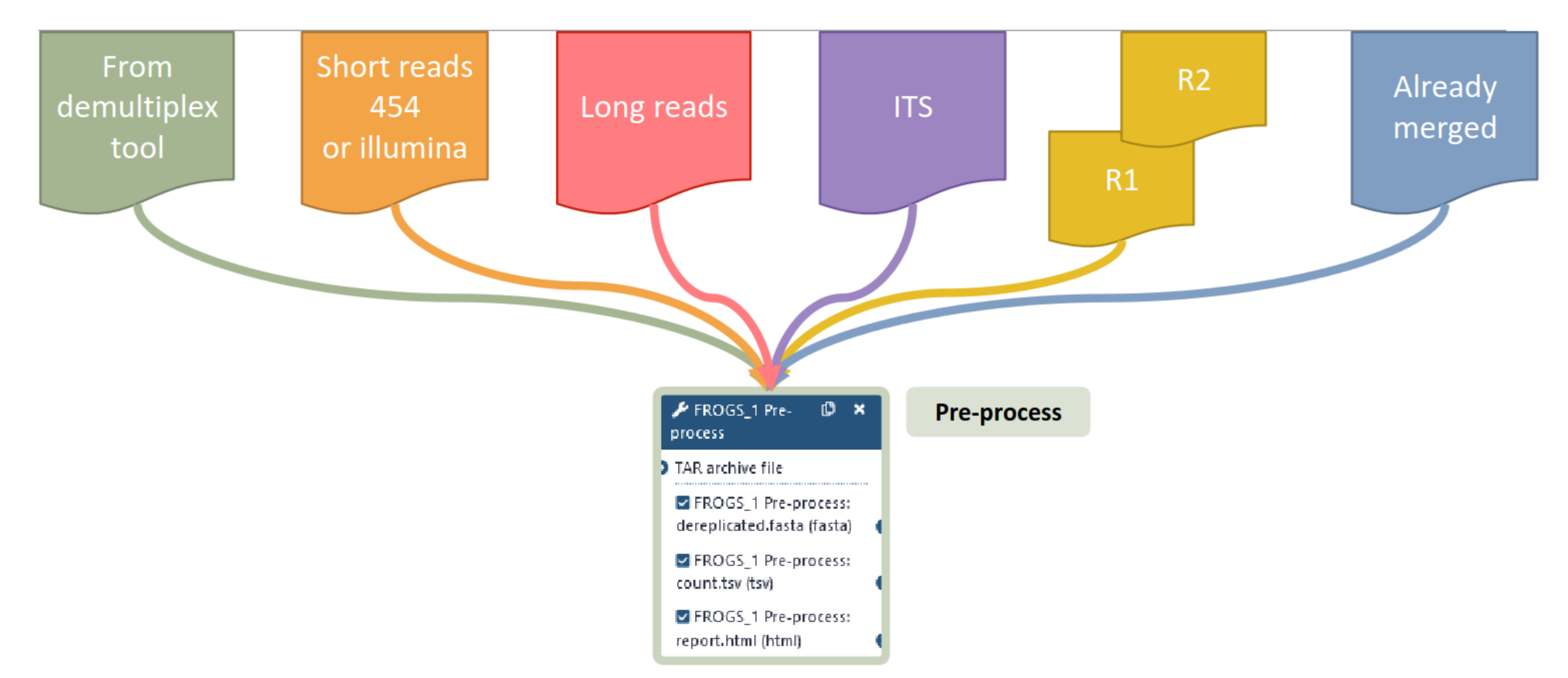
What FROGS preprocess does?
- Merging of R1 and R2 reads with vsearch [4], flash [5] or pear [6] (only in command line)
- Deletes sequences without good primers
- Finds and removes adapter sequences with cutadapt
- Deletes sequence with not expected lengths
- Deletes sequences with ambiguous bases (N)
- Dereplication
- removing homopolymers (size = 8) for 454 data
- quality filter for 454 data
Merging of paired-end reads
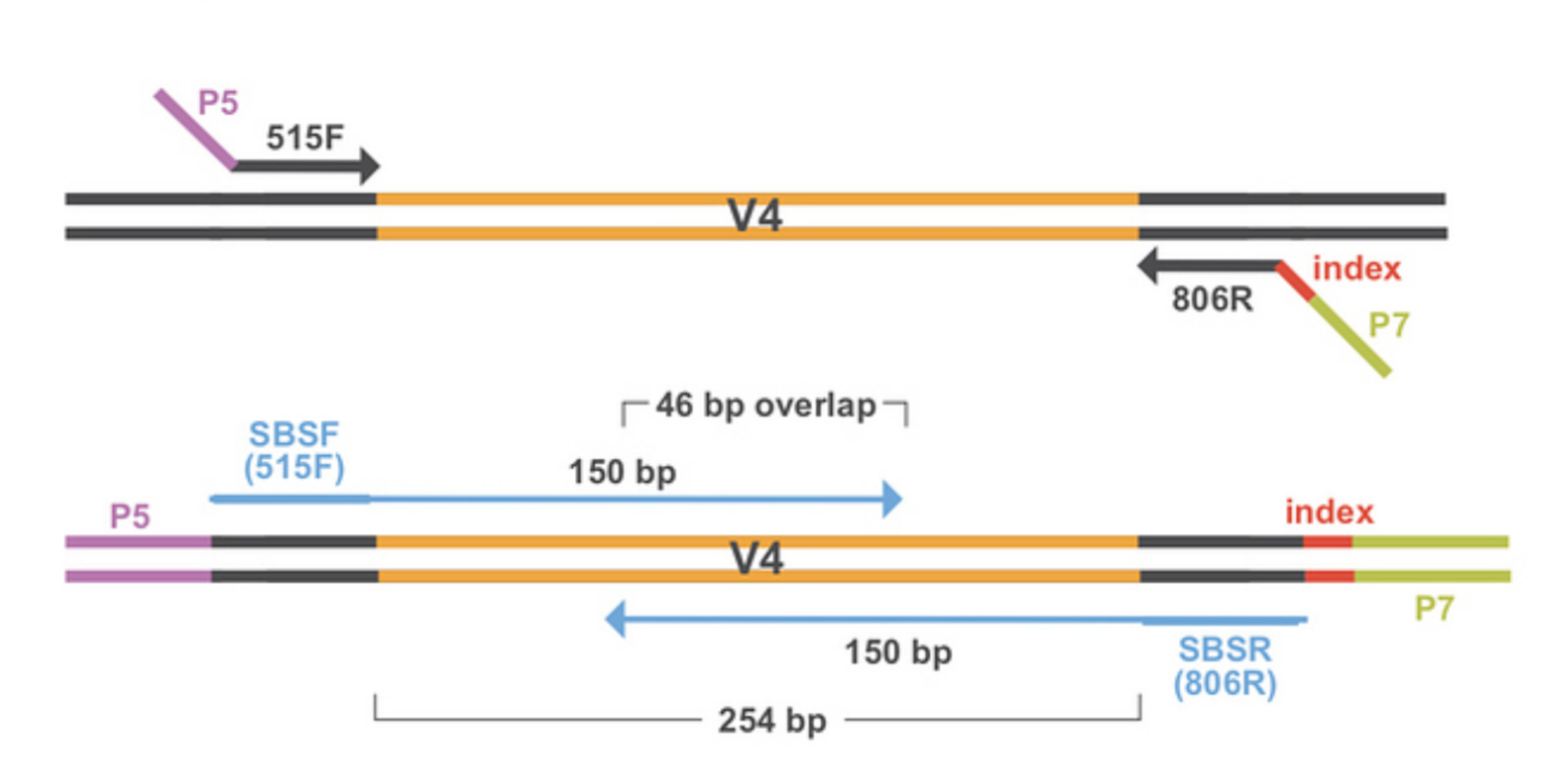
Clustering

Sequencing data are noised
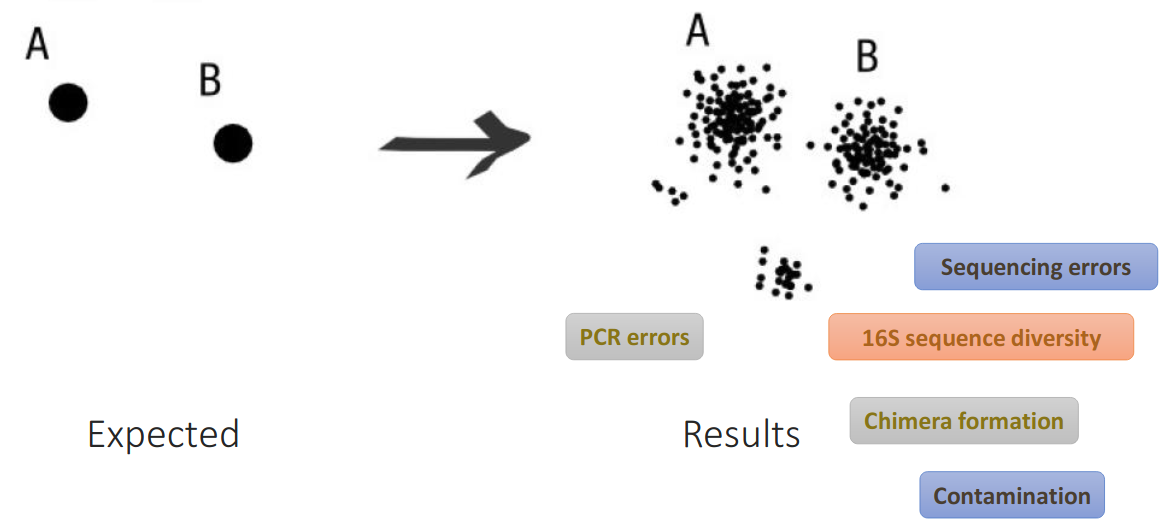

How to deal with these noised sequences?
- Comparison all against all
- Very accurate
- Requires a lot of memory and/or time
- Clustering
- closed-reference / open-reference
- de novo
- Denoising
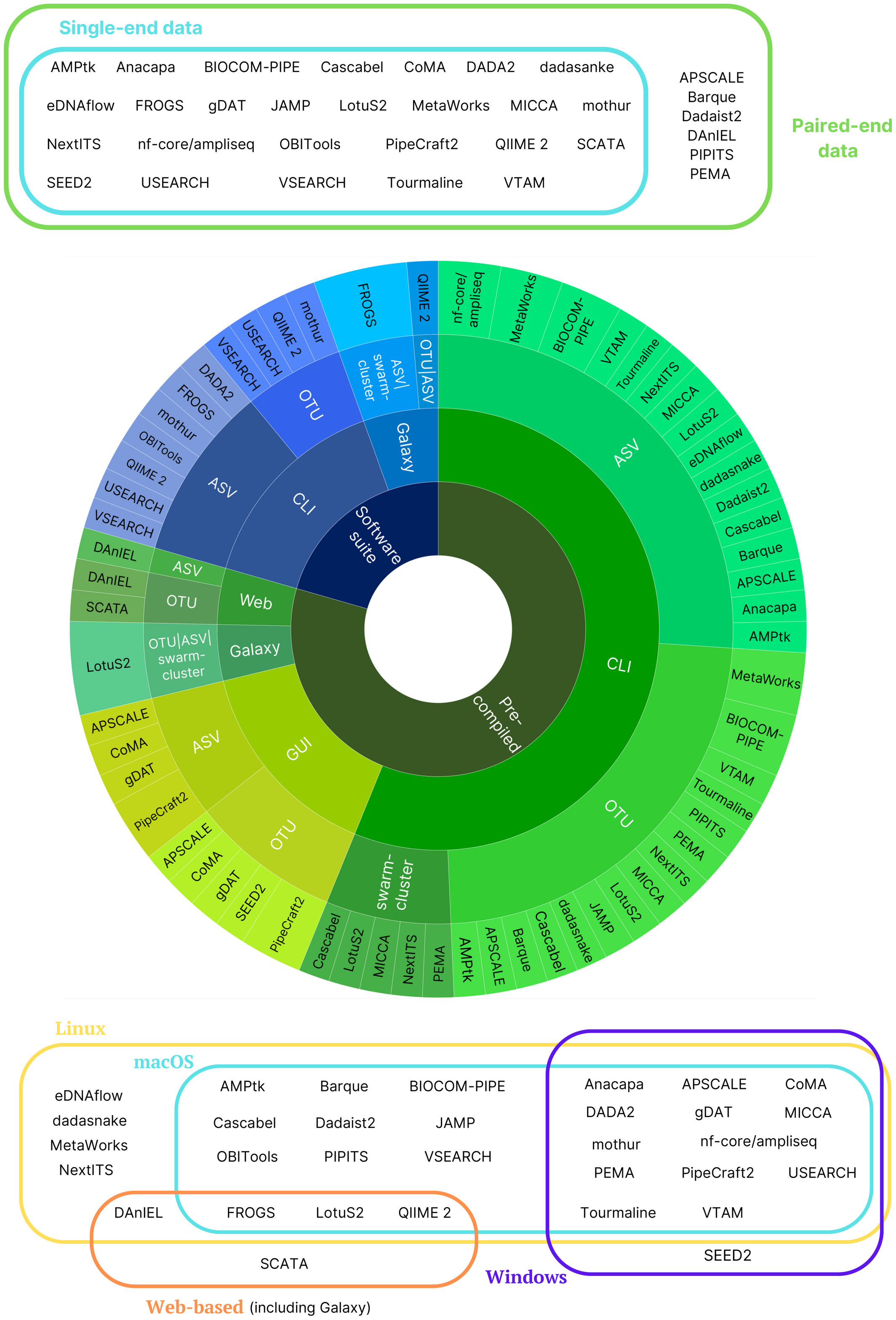
Methods classification
- A lot of terms for features built by softwares
- OTUs, zOTUS, ASVs, ESVs…
- A recent review establishes the vocabulary [7]
- OTUs / ASVs / swarm clusters
- ASVs are identical denoised reads with as few as 1 base pair difference between variants, representing an inference of the biological sequences prior to amplification and sequencing errors
- OTUs are formed with a % threshold clustering
- Swarm clusters are a third feature type
Methods classification
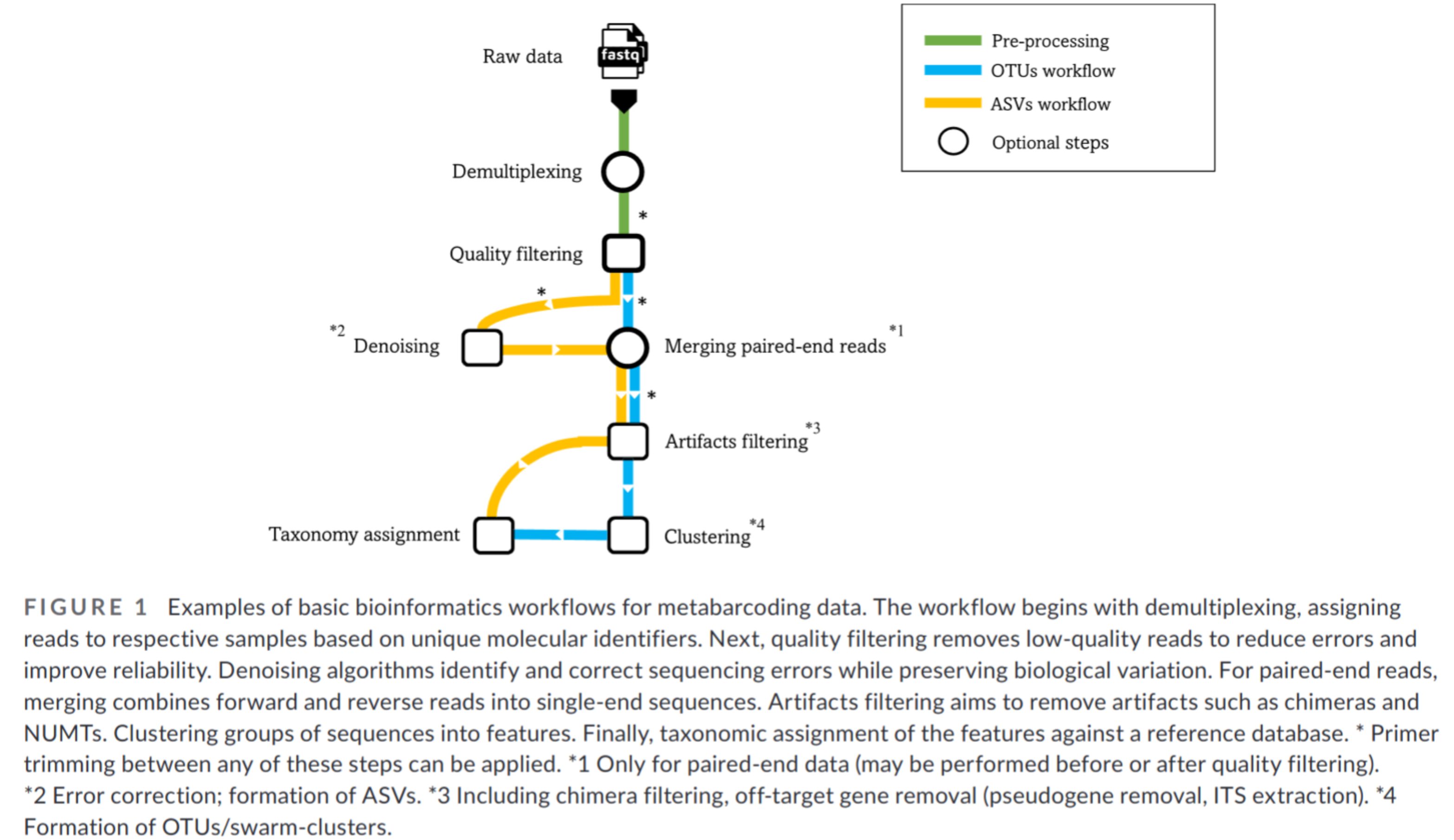
OTU paradigm
- Operational Taxonomic Unit
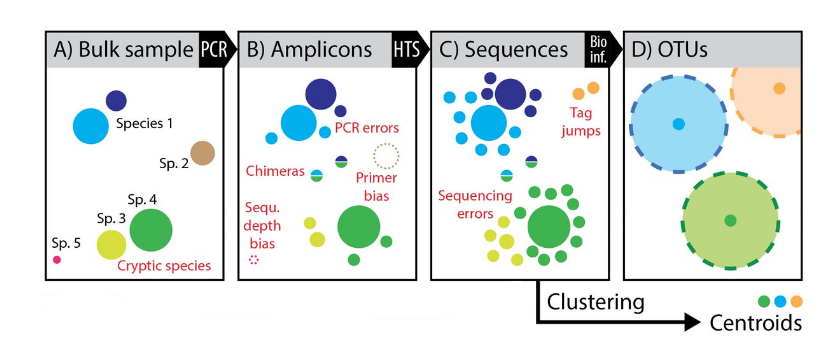
Operational Taxonomic Units
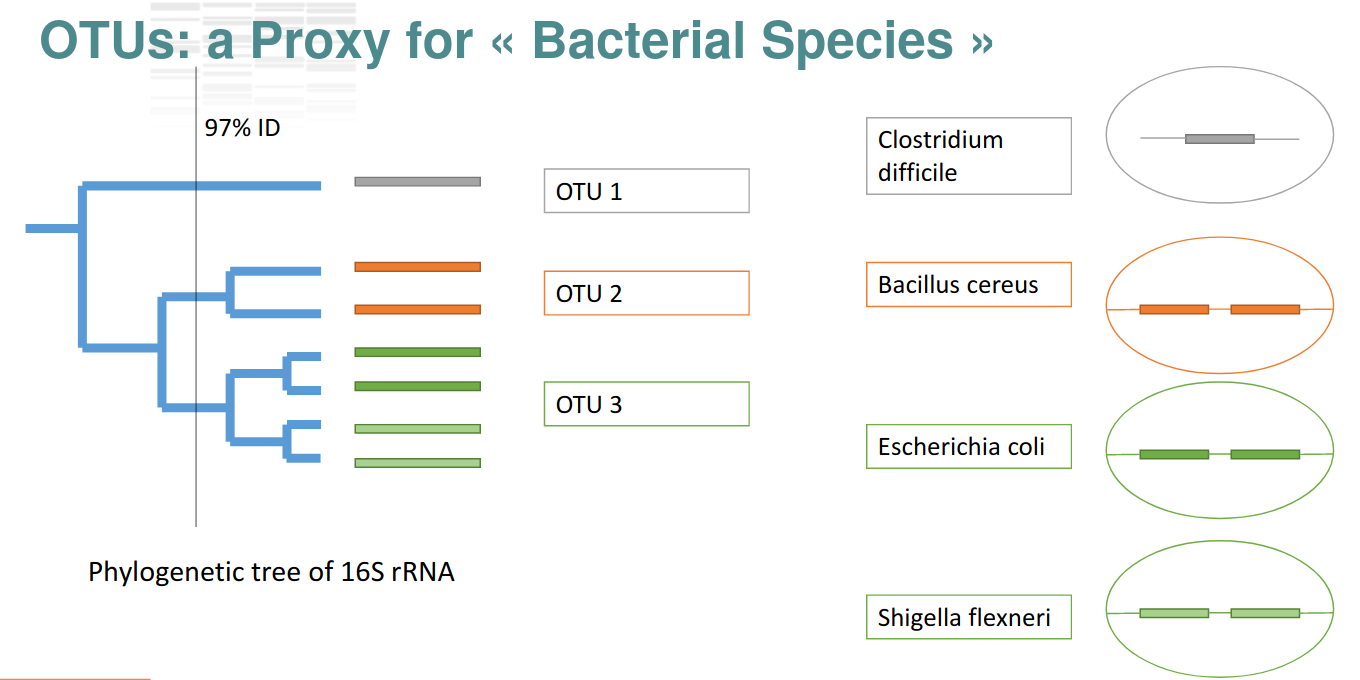

Operational Taxonomic Units
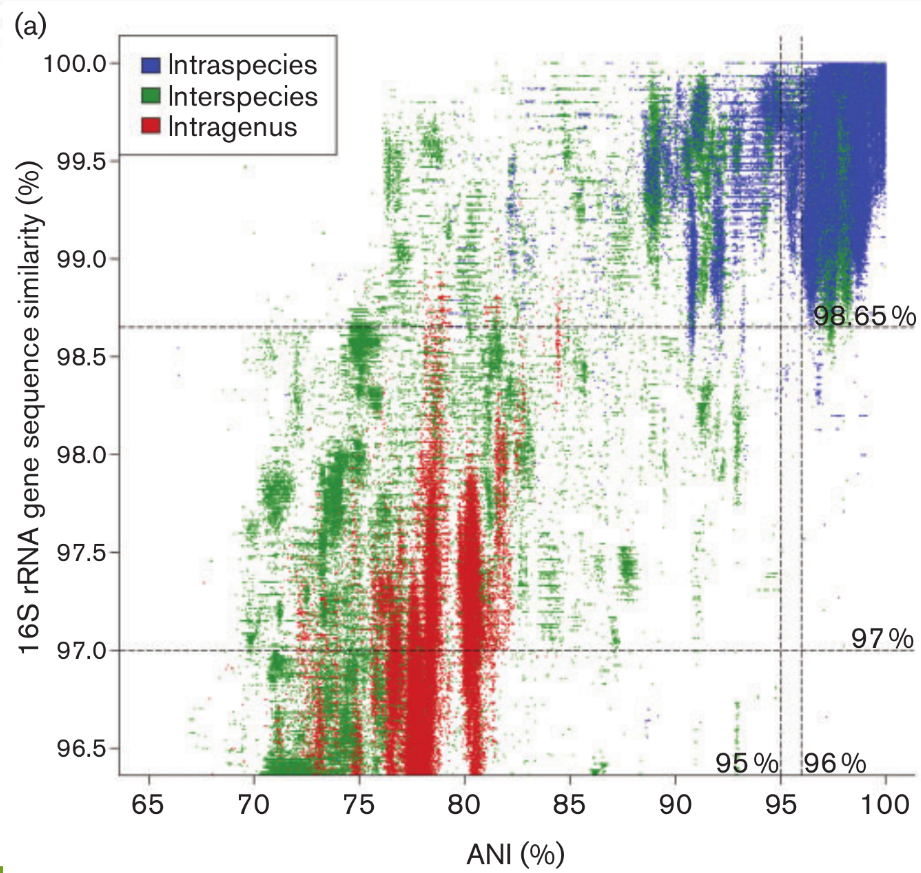
Operational Taxonomic Units
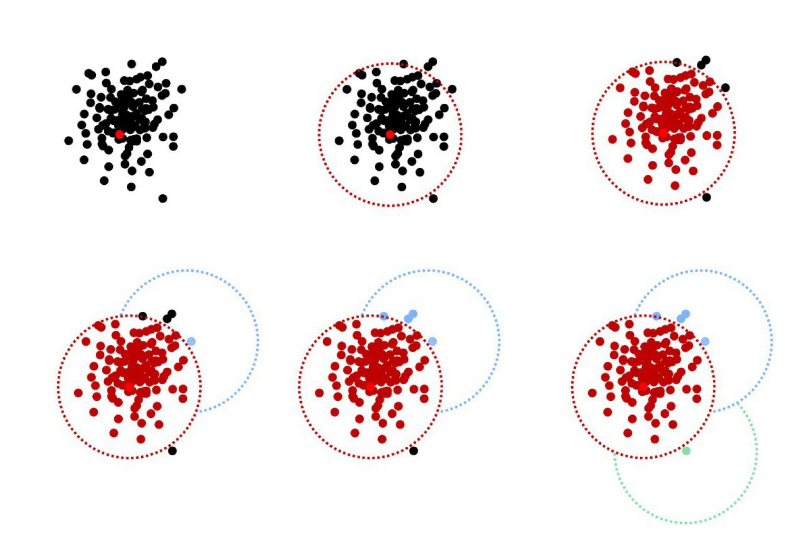
Operational Taxonomic Units
- OTUs composition is input-order depenent
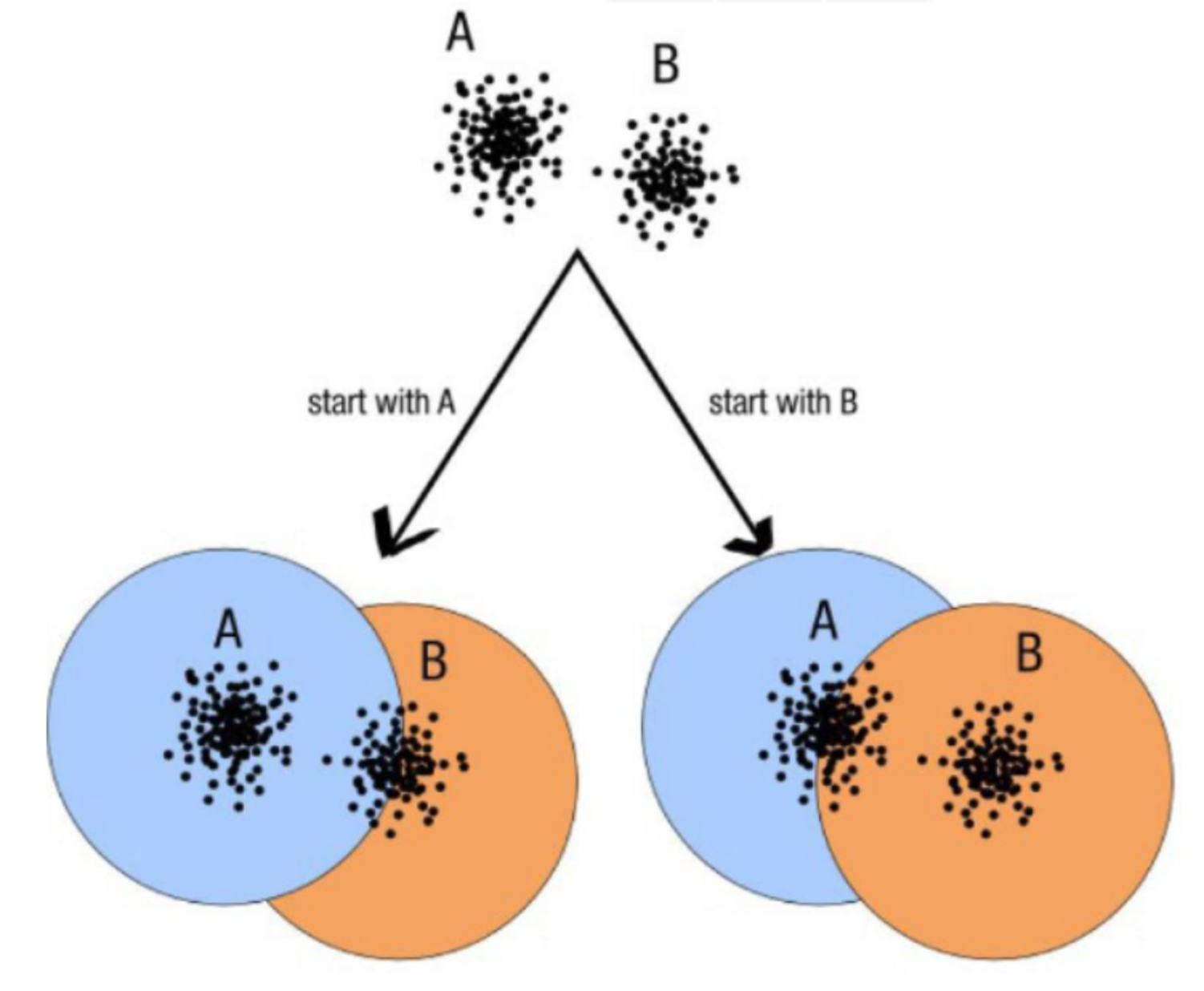
ASV paradigm
- Amplicon Sequence Variants
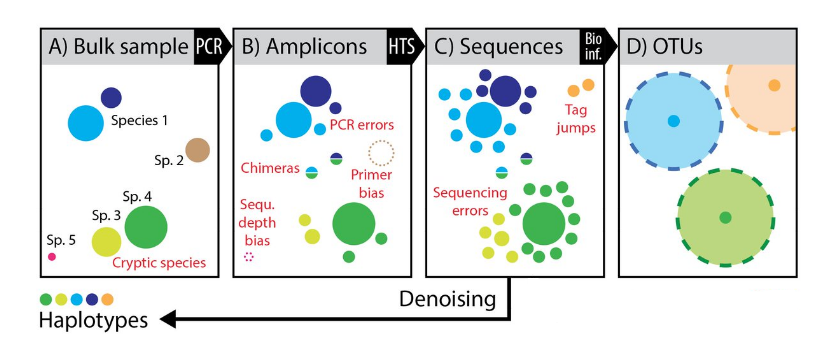
ASV are inferred by a de novo process in which biological sequences are discriminated from errors on the basis of the expectation that biological sequences are more likely to be repeatedly observed than are error-containing sequences
ASV resolution
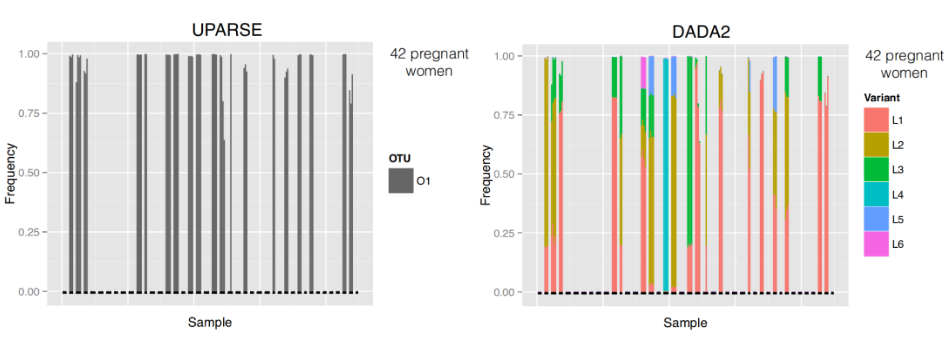
- ASV resolution changes the composition for these samples
Swarm
Swarm [8] is a notably different sequence clustering approach, which, while technically a clustering algorithm, may also be considered a denoising method when using the fastidious method with d=1. It relies on the maximum number of differences between reads (local linking threshold) and forms clusters that are resilient to input-order changes, thus creating stable, high-resolution features (herein referred to as swarm-clusters). When using the fastidious method with d=1, swarm aims to produce clusters centered around real biological sequences, where clusters represent sequence variants.
Since FROGS uses swarm (with the fastidious method with d=1) and strongly promotes denoising by chimera removal and cluster filtering, FROGS produces ASVs.
Why Swarm?
- Fixed clustering threshold is a real problem
- OTUs construction is input-order depenent
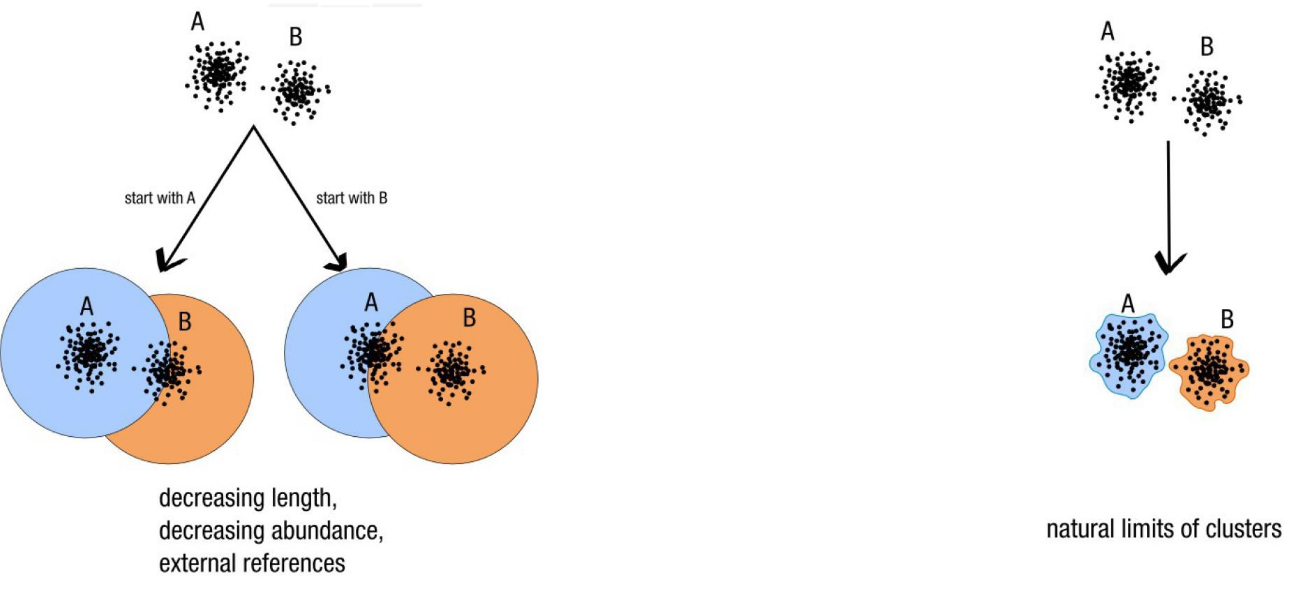
Swarm: A smart idea
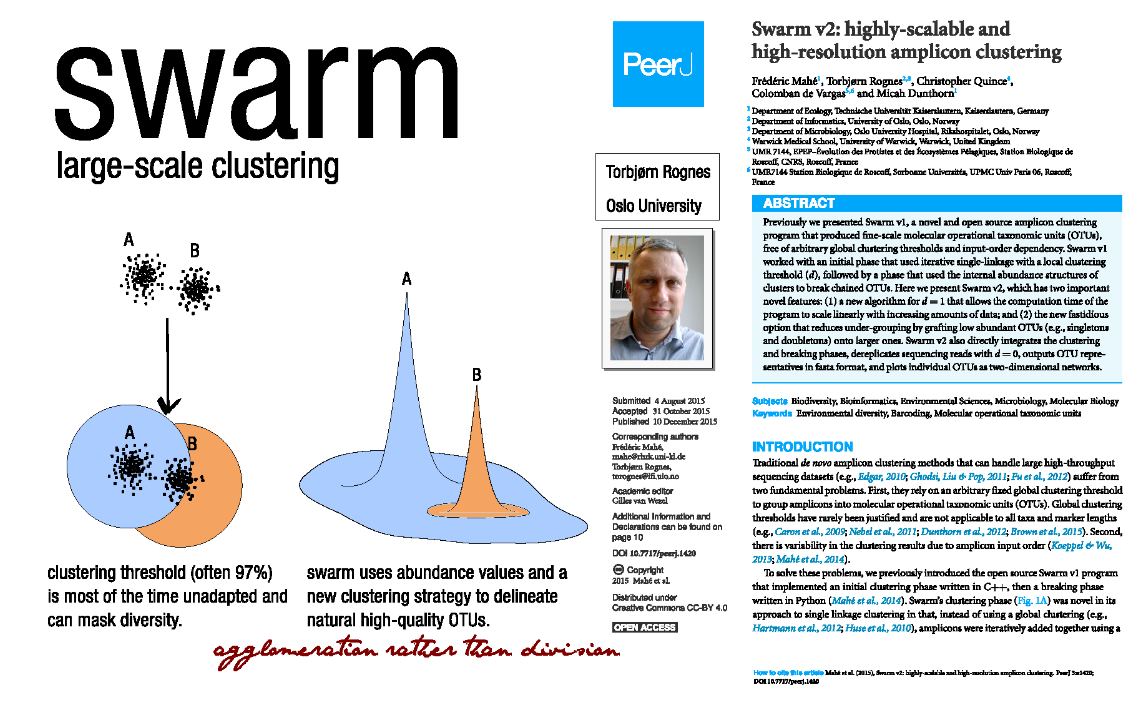
Swarm
- A robust and fast clustering method for amplicon-based studies
- The purpose of swarm is to provide a novel clustering algorithm to handle large sets of amplicons
- swarm results are resilient to input-order changes and rely on a small local linking threshold d, the maximum number of differences between two amplicons
- swarm forms stable high-resolution clusters, with a high yield of biological information
- Default: forms a lot of low-abundant OTUs that are in fact artifacts and need to be removed
- Swarm (fastidious method + d=1) clusters + filters → ASVs
d: the small local linking threshold
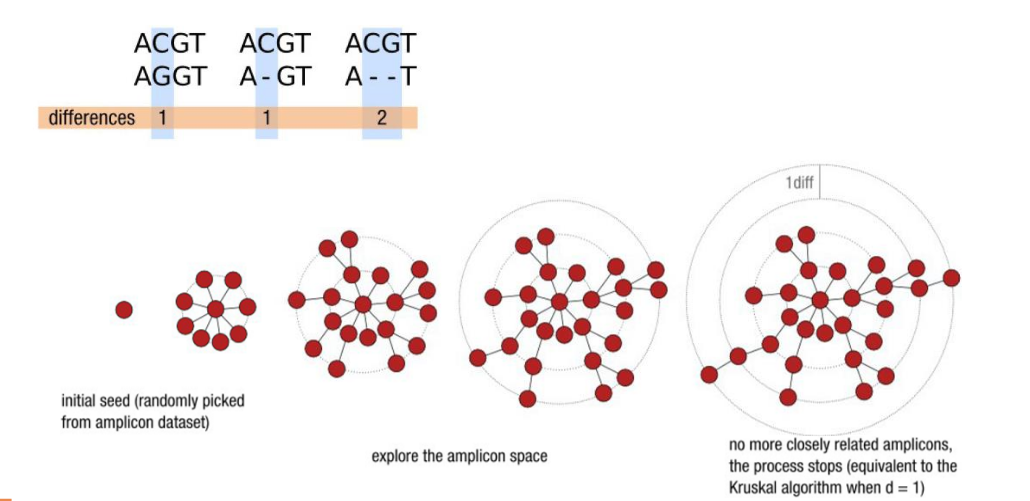
Swarm steps
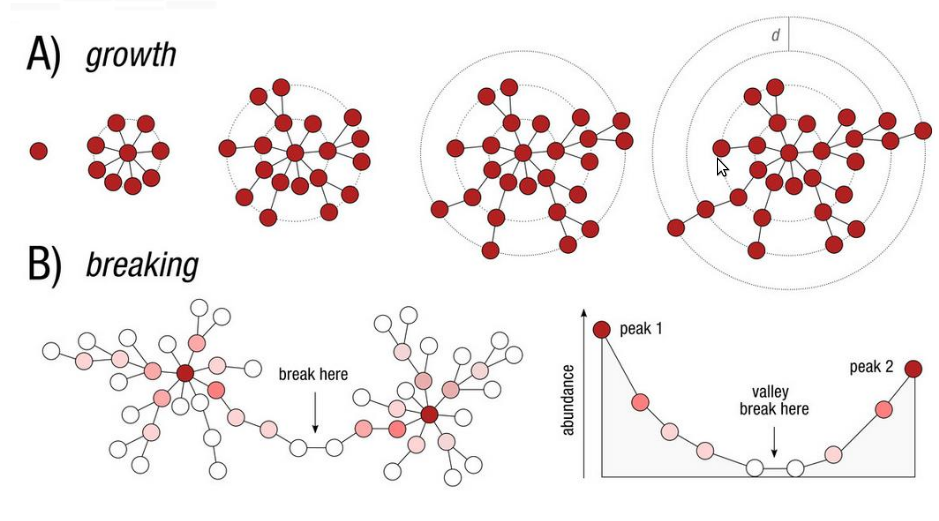
Which method to choose?

Advantages and inconvenients
- Which type of features to prefer may be context-dependent, and both may even be used in the same study
- ASVs demonstrate a biologically informative fine-scale resolution [9]
- But difficult to separate noise from a real signal in low abundant reads [10]
- ASVs represent stable and reproducible units across studies whereas OTUs are dataset-specific features (swarm clusters are not )
- problematic for longitudinal and very big studies
FROGS will soon offer the choice between swarm and dada2 for ASV creation
Chimera removal

Chimera removal


Chimera detection strategies
Reference based: against a database of «genuine» sequences
- dependant of the references used
De novo: against abundant sequences in the samples
FROGS uses vsearch [4] as chimera removal tool
A little extra: the sample-cross validation
- FROGS adds a sample-cross validation
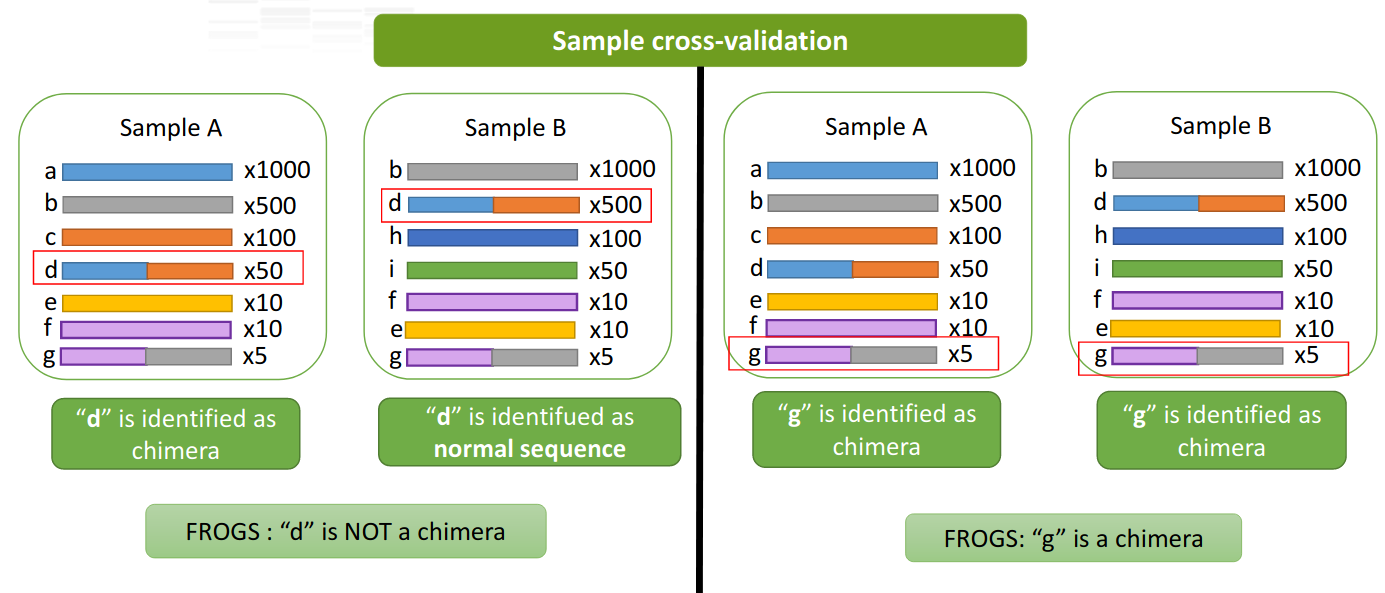
Chimera rates in samples
- From 5 to 40% in 16S data
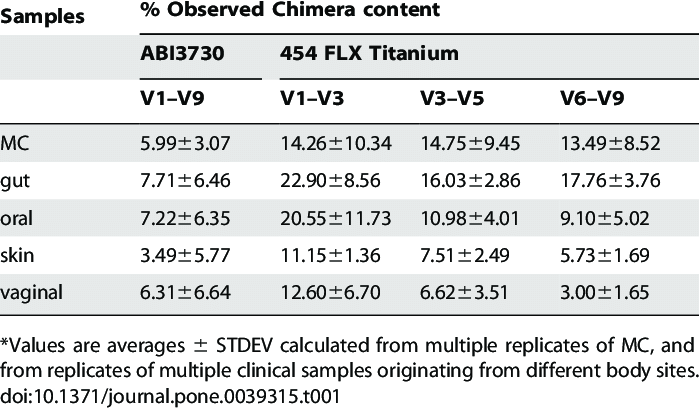
- Few with ITS (<10%)
Cluster filters

How to filter clusters?
- Low abundant sequences
- Clusters not shown in few replicates
- Contamination
Taxonomic affiliation

Comparison of approaches
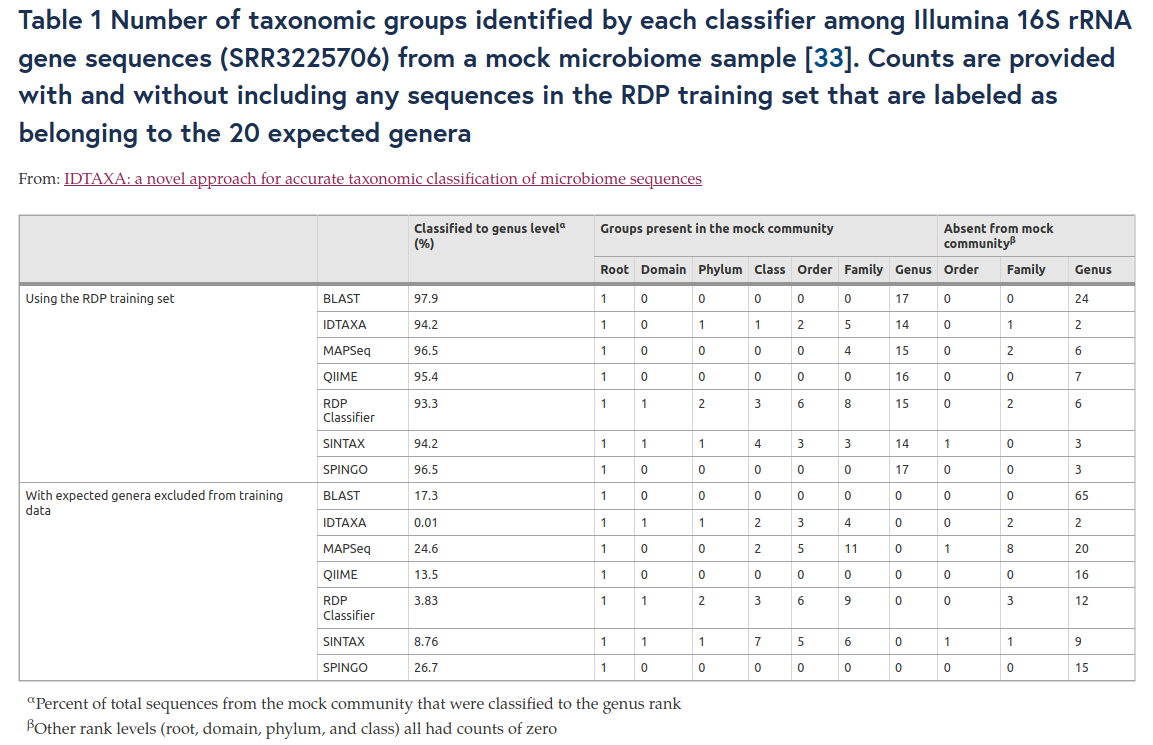
RDP problems
- Depends too much on the databank used!
- Gives one affiliation for each feature with bootstrap, on each subdivision
Bacteria;(1.0);
Actinobacteriota;(1.0);
Actinobacteria;(1.0);
Propionibacteriales;(1.0);
Propionibacteriaceae;(1.0);
Cutibacterium;(1.0);
Cutibacterium acnes;(0.57);The FROGS recommandation
- Use Blast and not RDP
- Check Blast metrics to avoid concluding too fast
- Take care of the reference databank used!
Bacteria;
Actinobacteriota;
Actinobacteria;
Propionibacteriales;
Propionibacteriaceae;
Cutibacterium;
Multi-AffiliationThe FROGS databanks
- Command line: you can use your own databank
- Galaxy
- You have access to several databanks
- Admins have to add your databank
- The file must be well formated, we can do it for you
- For private databanks, contact us!
The FROGS extra: the multi-affiliations
- FROGS gives all identical hits
Bacteria;Firmicutes;Bacilli;Staphylococcales;Staphylococcaceae;Staphylococcus;
Staphylococcus xylosus
Bacteria;Firmicutes;Bacilli;Staphylococcales;Staphylococcaceae;Staphylococcus;
Staphylococcus saprophyticusStrictly identical (V1-V3 amplification) on 499 nucleotides
- FROGS can’t decide if it’s one or another
- You have to check if you can choose between multi-affiliations
To help you
- https://shiny.migale.inrae.fr/app/affiliationexplorer
- a very user-friendly Shiny web app, allowing users to modify very simply the affiliations from a FROGS abundance file
Demo
Filter ASVs on their affiliation

Affiliation filters
Remaining contamination?
Want to analyse only the
Firmicutes?2 modes
- Deleting: remove ASVs
- Hiding: only the affiliation is modified, not the abundance
Phylogenetic tree
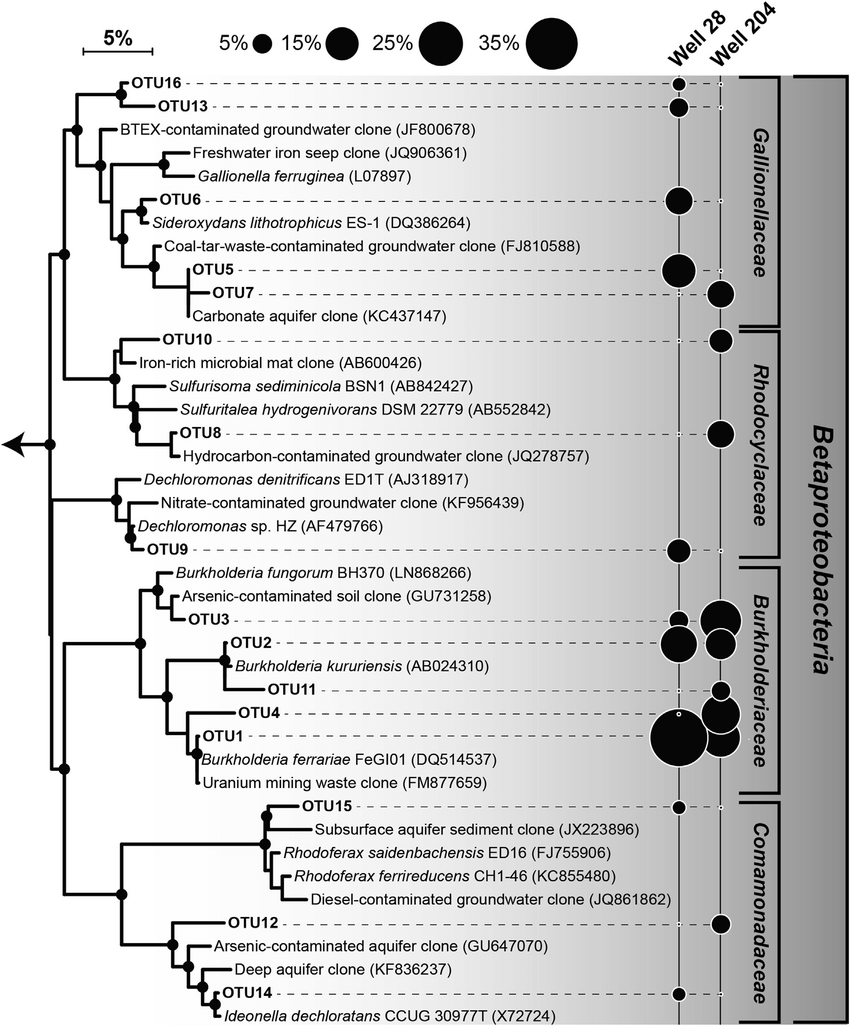
FROGS tree
- This tool builds a phylogenetic tree thanks to affiliations of ASVs contained in the BIOM file
- Needed to compute beta-diversity indices based on phylogenetic distances
- Interesting to explore poor-characterized environments
FROGSfunc: function inference
Concepts
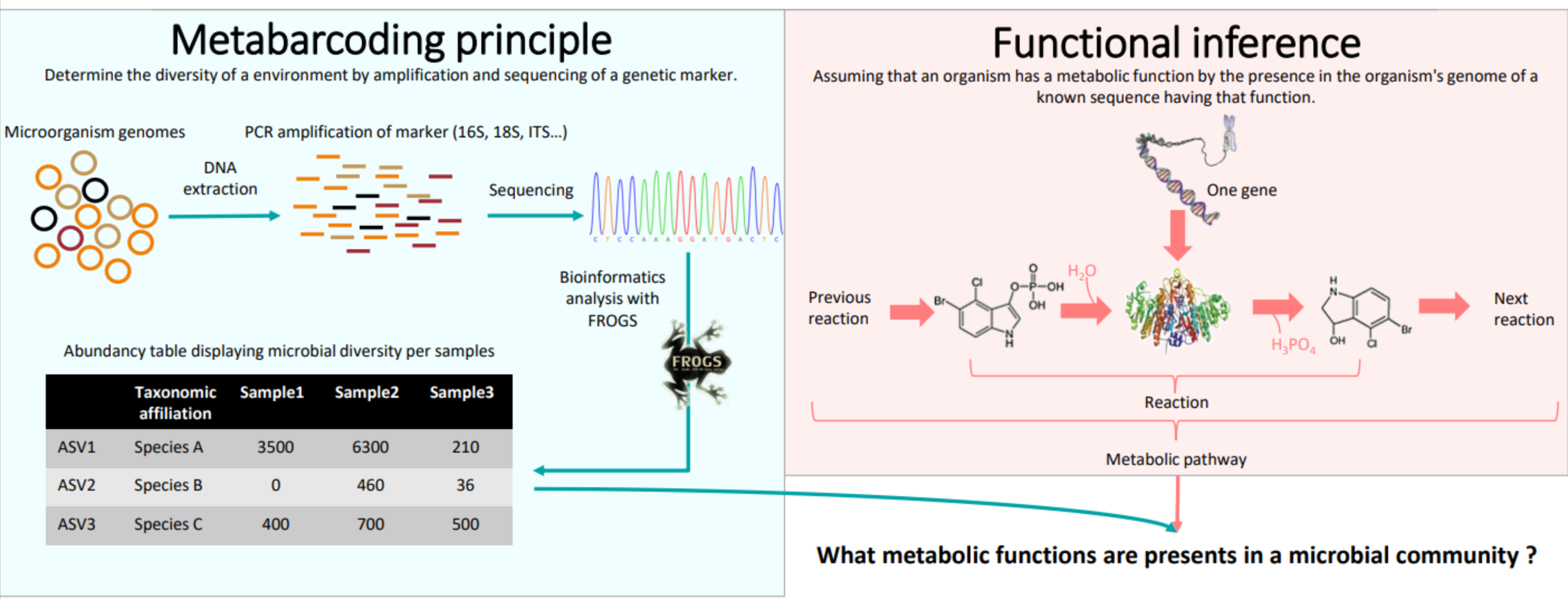
Based on PICRUSt2
- PICRUSt [13] (Phylogenetic investigation of communities by reconstruction of unobserved states) is an open-source tool.
- It is a software for predicting functional abundances based only on marker gene sequences
- PICRUSt2 is composed of 4 python applications.
- No graphic interface exists to run PICRUSt2 for non-expert users.
How it works
- Places the ASVs into a reference phylogenetic tree and predicts of marker copy number in each ASV.
- Predicts number of function copy number in each ASV and calculates functions abundances in each sample and ASV abundances according to marker copy number.
- Calculates pathway abundances in each sample.
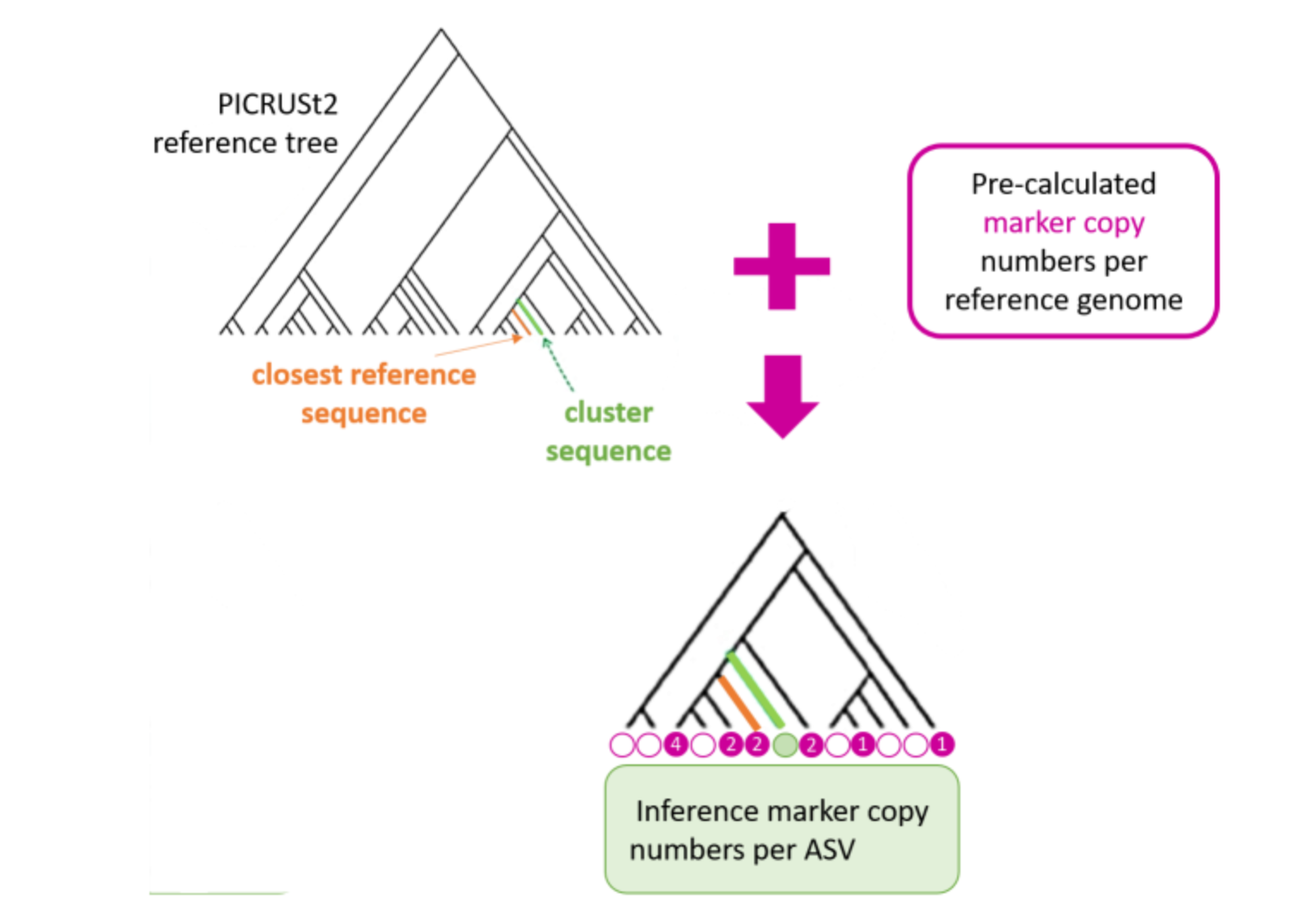
FROGSfunc placeseqs and copynumber
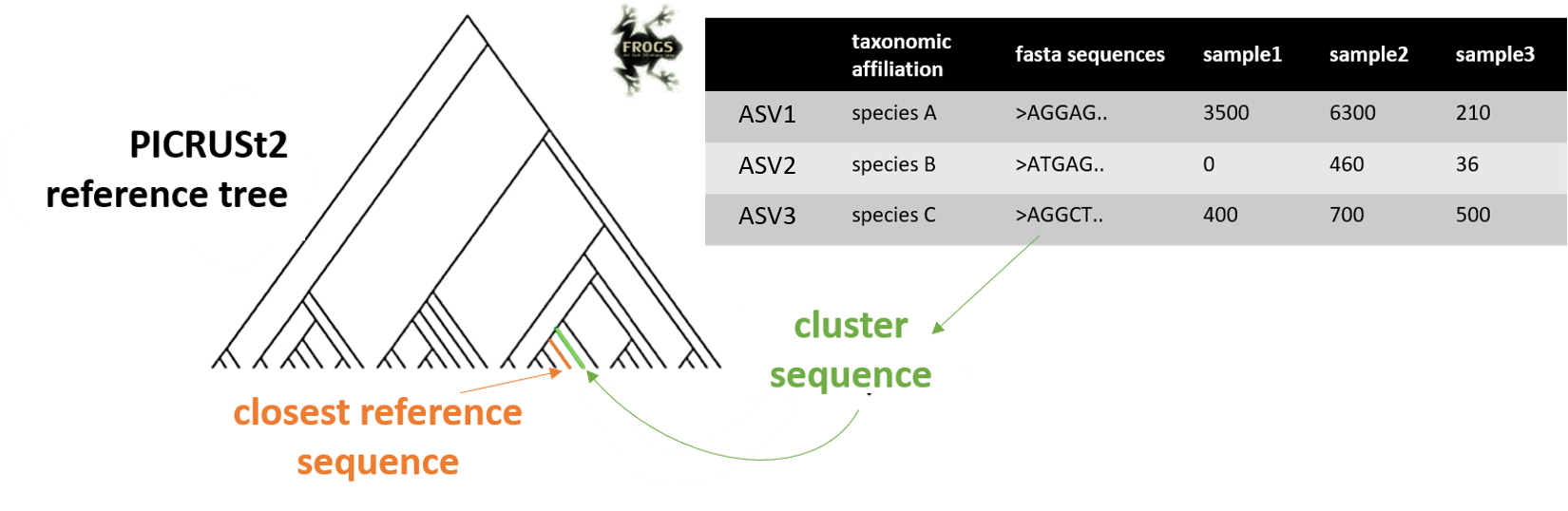
FROGSfunc placeseqs and copynumber
NSTI
- NSTI scores are simply the average branch length that separates each ASV in your sample from a reference bacterial genome, weighted by the abundance of that ASV in the sample.
- PICRUSt2 sets NSTI threshold to 2 per default. Some studies have shown that this threshold is permissive. Thus, it is important to see if the taxonomies between PICRUSt2 and FROGS are quite similar or not, in order to potentially choose a more stringent threshold afterwards.
- 0 < Good < 0.5
- 0.5 <= Medium < 1
- 1 <= Bad < 2
- To exclude >= 2
FROGSfunc functions
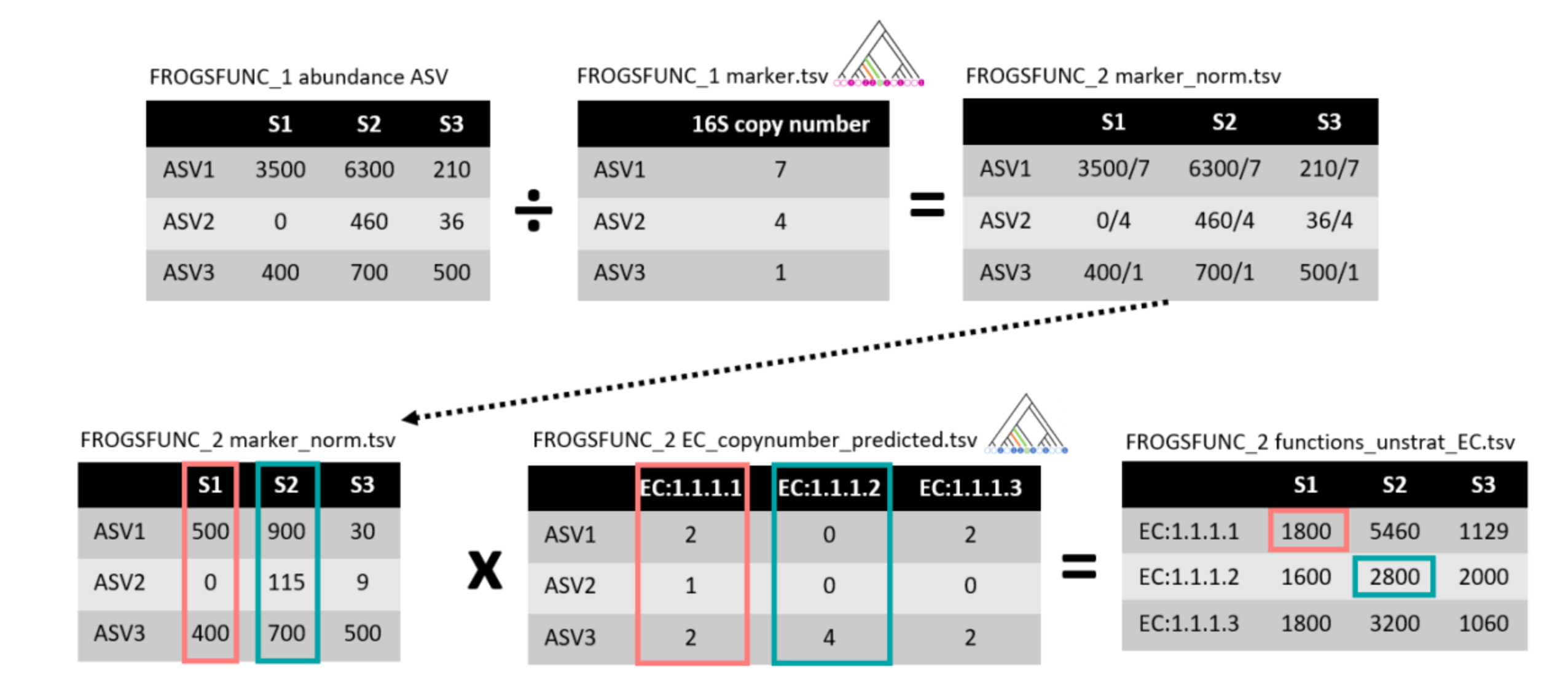
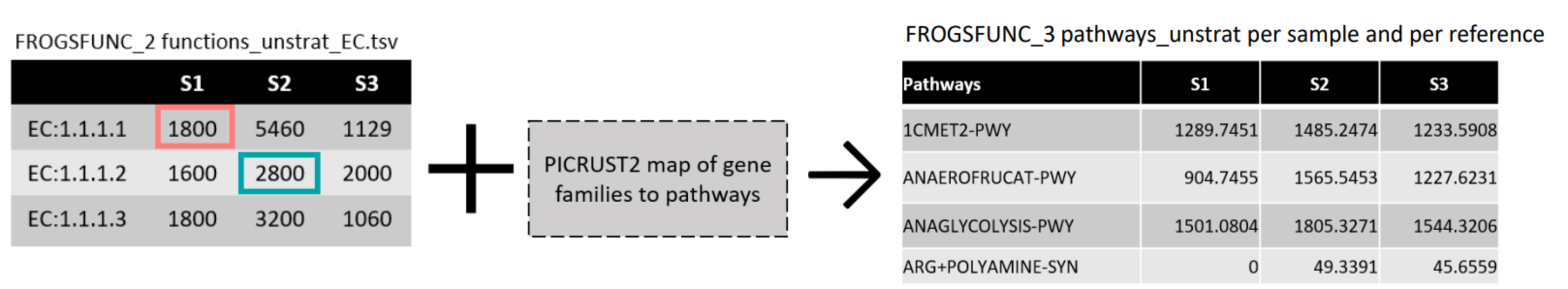
FROGSfunc pathways
Thanks for your attention

References

Master BMC - Atelier Biodiversité microbienne
