Comparison of metabarcoding taxonomic markers to describe fungal communities in fermented foods
METABARFOOD project
December 4, 2023
Introduction
Meta-omics using next-genertation sequencing (NGS)
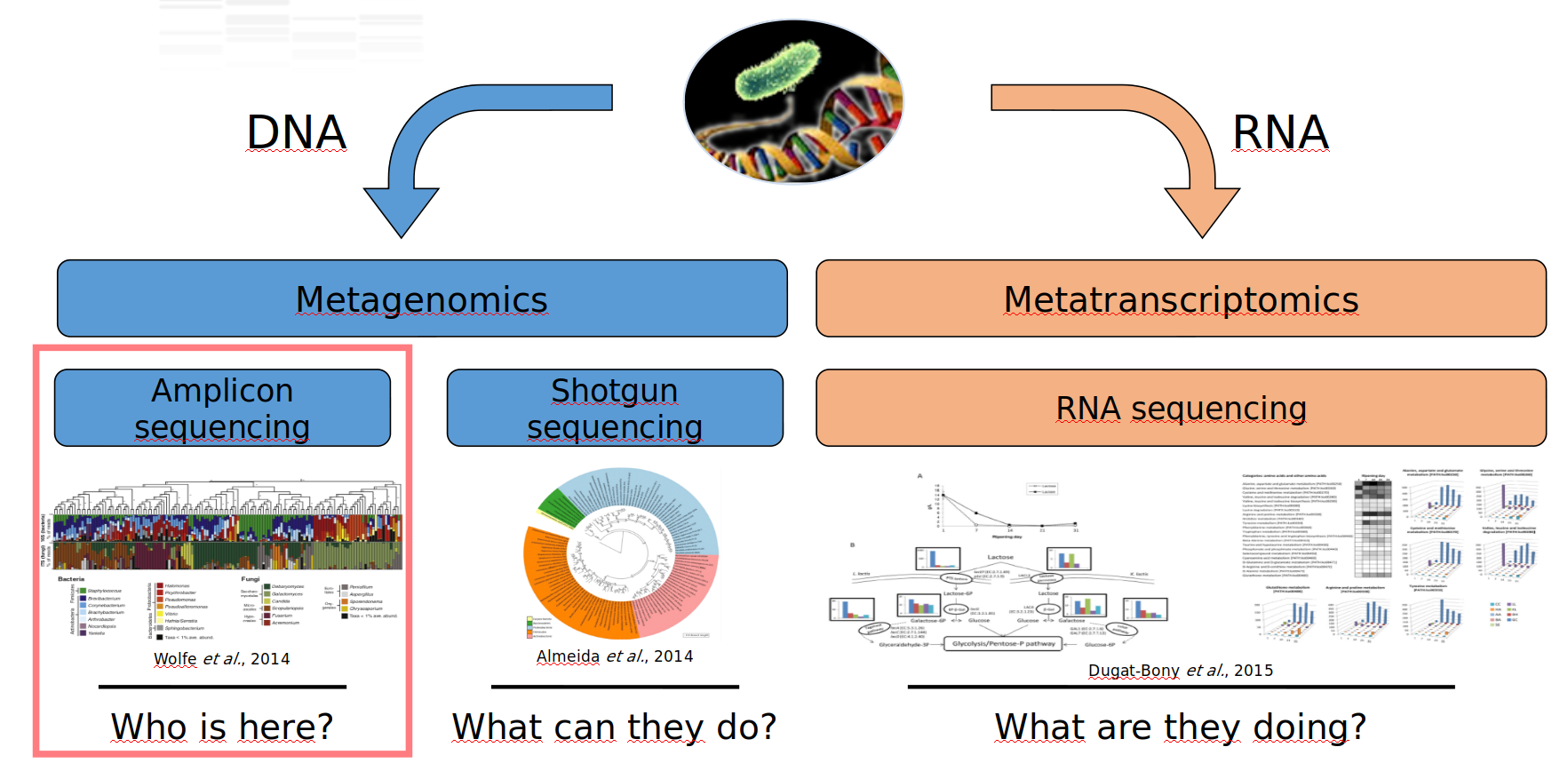
Meta-omics using next-genertation sequencing (NGS)
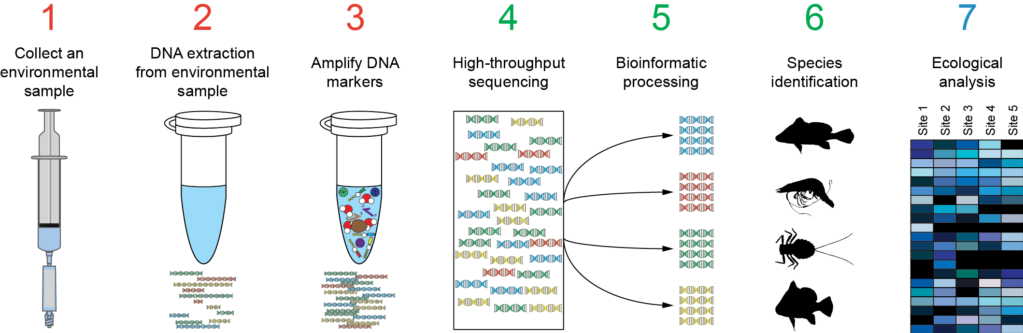
Metabarcoding
-
Advantages
- Analyze and compare many taxa (hundreds) at the same time
- Access to uncultured organisms
- Taxonomic profiles of the communities
- Easy to analyze compared to metagenomics
-
Weaknesses
- Many biases (amplification, extraction, sequencing…)
- Exact identification of the organisms difficult or impossible
- No functional view of the ecosystem
LOW COST: <35$ per sample
What is a good marker gene?
- ubiquist, conserved among taxa
- enough divergent to distinguish strains, not submitted to lateral transfer
- only one copy in genome
- has conserved regions to design specific primers
- is enough studied to be present in databanks for taxonomic affiliation
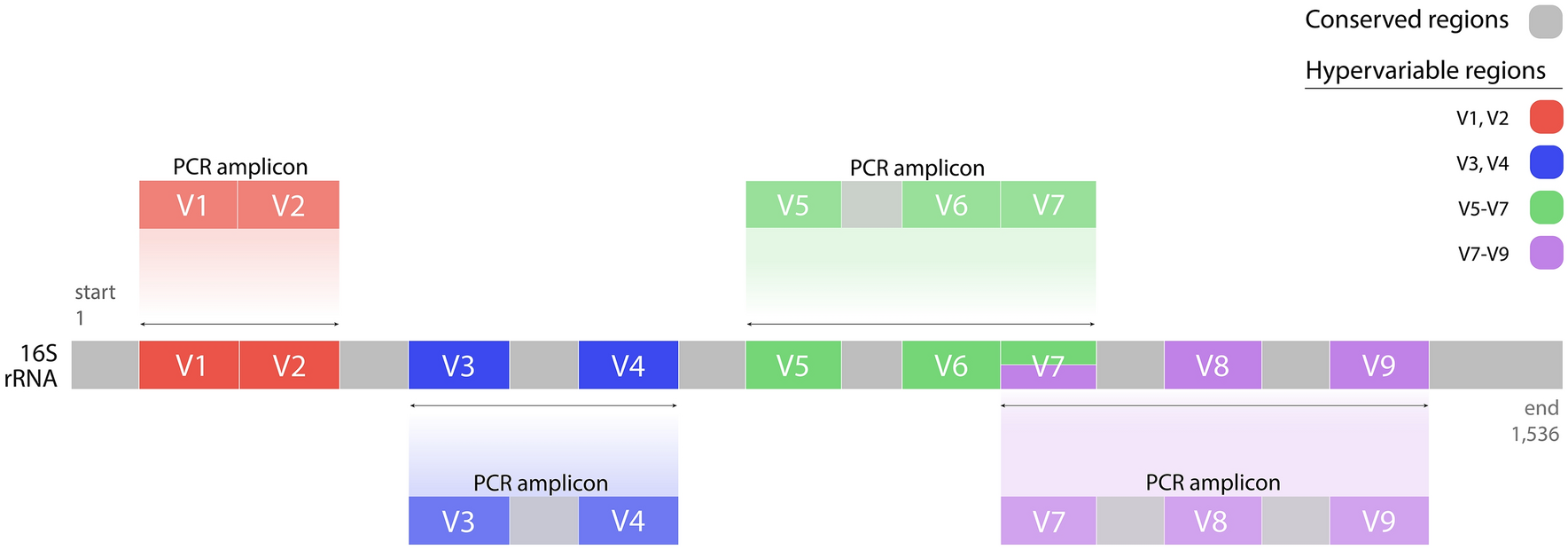
16S for bacteria, 18S for eukaryota, COI for plants… and for fungi?
- The Fungal Barcoding Consortium recommended the use of the Internal Transcribed Spacer (ITS) region as the primary marker for fungal identifications due to superior species-level resolution compared to LSU and SSU rRNA genes
- variable ITS length: up to 800/1000 bp ⚠️
- However, amplicon-based metagenetic analysis using MiSeq sequencing imposes more constraints than classical species identification. Most importantly, the typical amplicon size (~500bp) will not provide complete ITS region sequences or full length rRNA or protein coding genes
- The most popular barcodes currently used for fungal community analysis by metabarcoding are ITS1, ITS2 and the LSU D1/D2 domain
- variable amplicon length: 50 to 1200 bp ⚠️
Size and Illumina sequencing
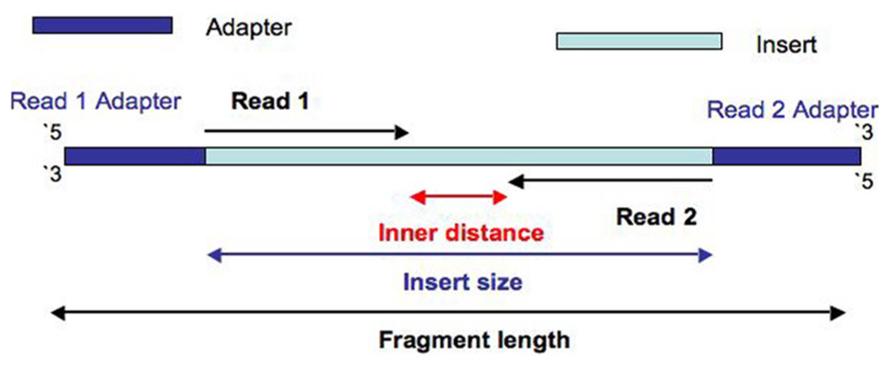
Bioinformatics challenges
- ⚠️ No tool able to deal correctly with sequences of variable sizes → impossible to compare them
- All tools dedicated to 16S data (350-500 nt)
- Adaptation of FROGS [1] to deal with ITS data [2]


Management of short fragments
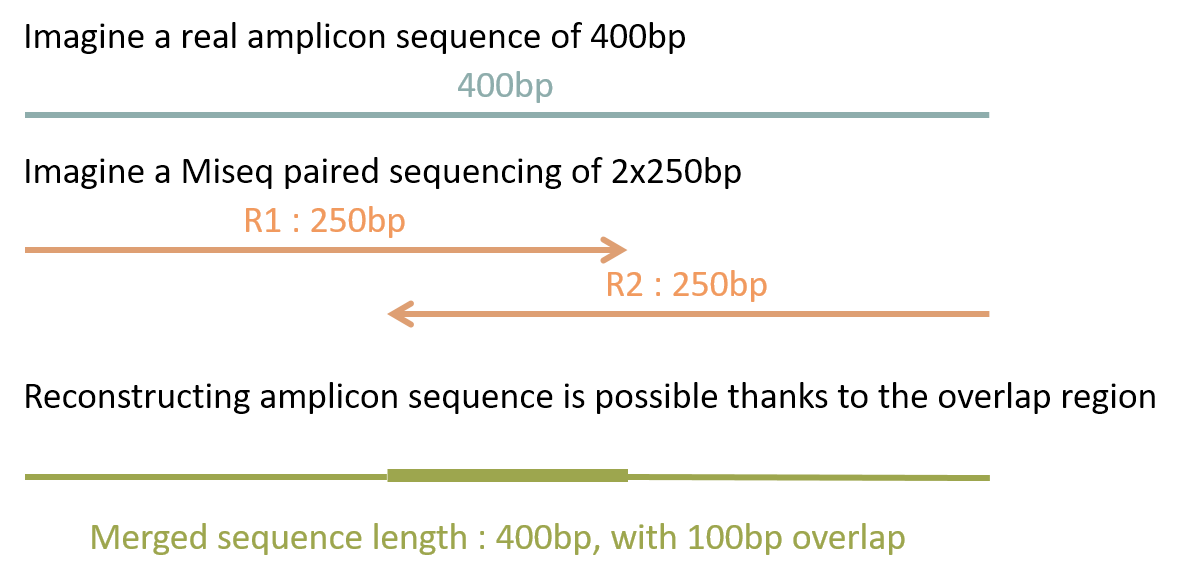
| FROGS | Qiime2 | DADA2 | USEARCH | |
|---|---|---|---|---|
| Short | ✅ | ✅ | ✅ | ✅ |
Management of long fragments
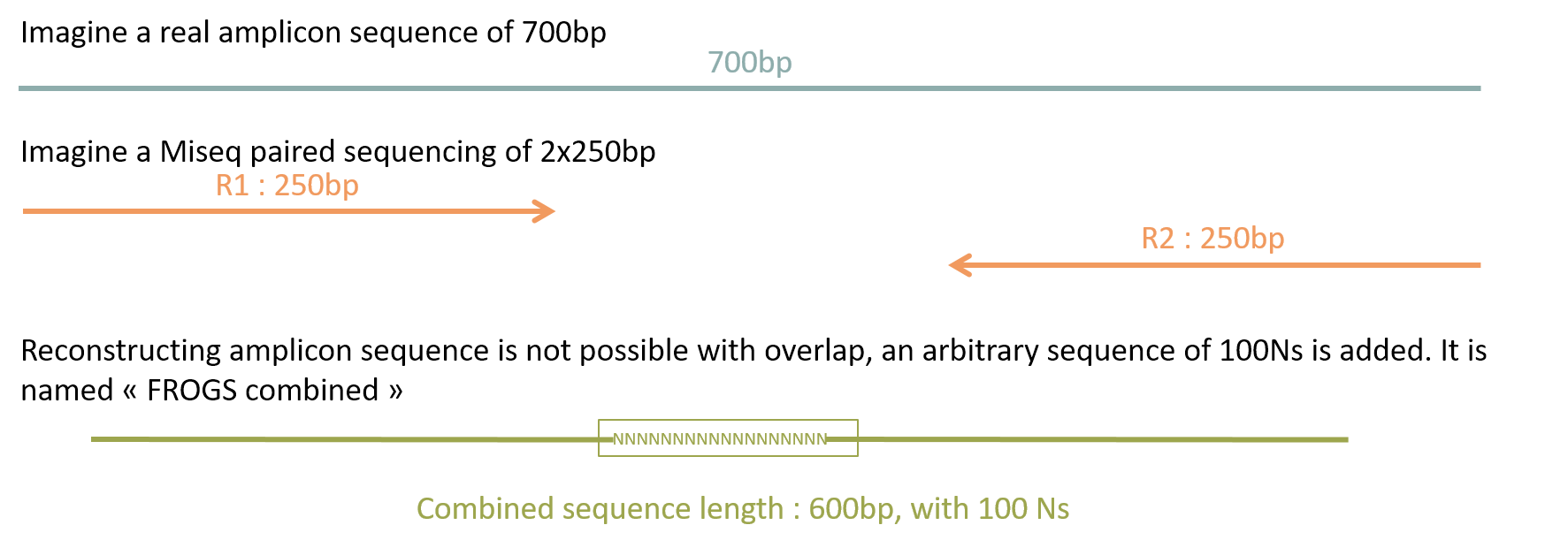
| FROGS | Qiime2 | DADA2 | USEARCH | |
|---|---|---|---|---|
| Long | ✅ | ❎ R1 only | ❎ R1 only | ❎ R1 only |
Management of mixed fragments
| FROGS | Qiime2 | DADA2 | USEARCH | |
|---|---|---|---|---|
| Mixed | ✅ | ❎ R1 or only short | ❎ R1 or only short | ❎ R1 and only short |

Clustering or denoising?
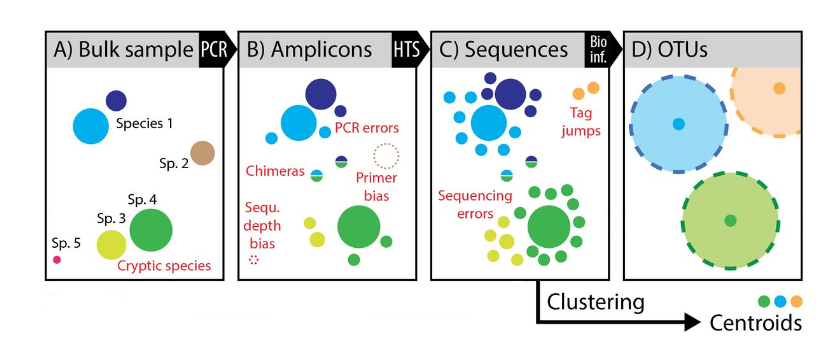
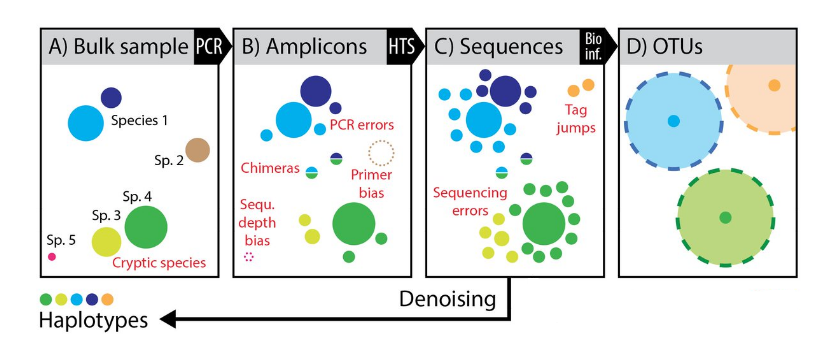
Densoising power
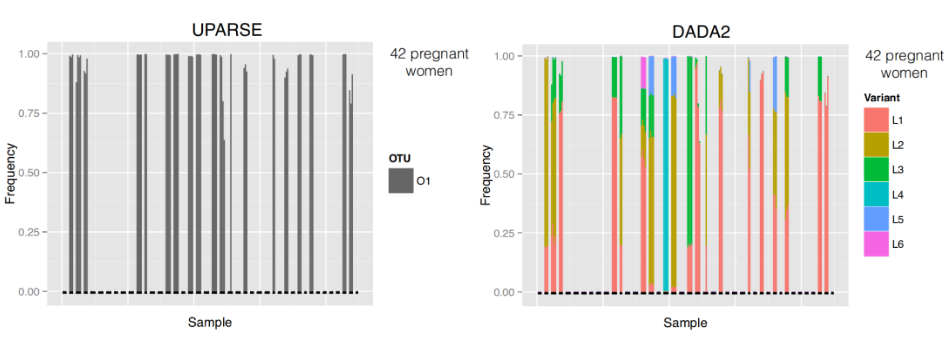
- Variants are aggregated in a single OTU
- Conclusions are not the same
Clustering or denoising?
| FROGS | Qiime2 | DADA2 | USEARCH | |
|---|---|---|---|---|
| Method | Swarm clusters (ASV) | Clustering 97% (OTU) | Denoising (ASV) | Denoising (zOTU) |
| Short fragments | ✅ | ✅ | ✅ | ✅ |
| Long fragments | ✅ | ❎ R1 only | ❎ R1 only | ❎ R1 only |
| Mixed fragments | ✅ | ❎ R1 or only short | ❎ R1 or only short | ❎ R1 and only short |
- FROGS only able to treat all data but…
- … denoising tools able to detect haplotypes
- … and are able to build consistent features among studies
How to reconcile the advantages of each side?
The METABARFOOD project (2015)
The METABARFOOD project (2015-2023)
- Project “Métaprogramme MEM” (2015) driven by Delphine Sicard (SPO)
-
Partners
- LUBEM (INRAE / Université Brest), SPO (INRAE Montpellier), SayFood (INRAE Saclay), Migale (INRAE Jouy-en-Josas)
-
Aims
- Standardize library prep
- Compare markers (ITS, D1/D2, RPB2) and metabarcoding pipelines to determine the fungal diversity in several food matrices (bread, fermented meat, wine, cheese)
- Test these approaches on real and mock communities
The METABARFOOD project (2015-2023)
-
Samples
- 96 samples from mock communities (27-60 species): 4 matrices x 4 markers x 2 conditions (PCR/DNA) x 3 replicates
- 144 real samples: 4 matrices x 4 markers x 3 samples x 3 replicates
- Illumina Miseq 2x250 sequencing (GeT-PlaGe Toulouse)
- 7 tools to compare
Note
Interesting challenge in the management of the samples and results
Methods
- For each fermented food type, representative species were selected based on an inventory of the most frequently described species in the literature. One strain (mainly available type-strains) was included for each selected species.
| Bread | Cheese | Meat | Wine | |
|---|---|---|---|---|
| Species | 27 | 25 | 40 | 60 |
| Genus | 15 | 16 | 14 | 37 |
| Family | 4 | 11 | 8 | 8 |
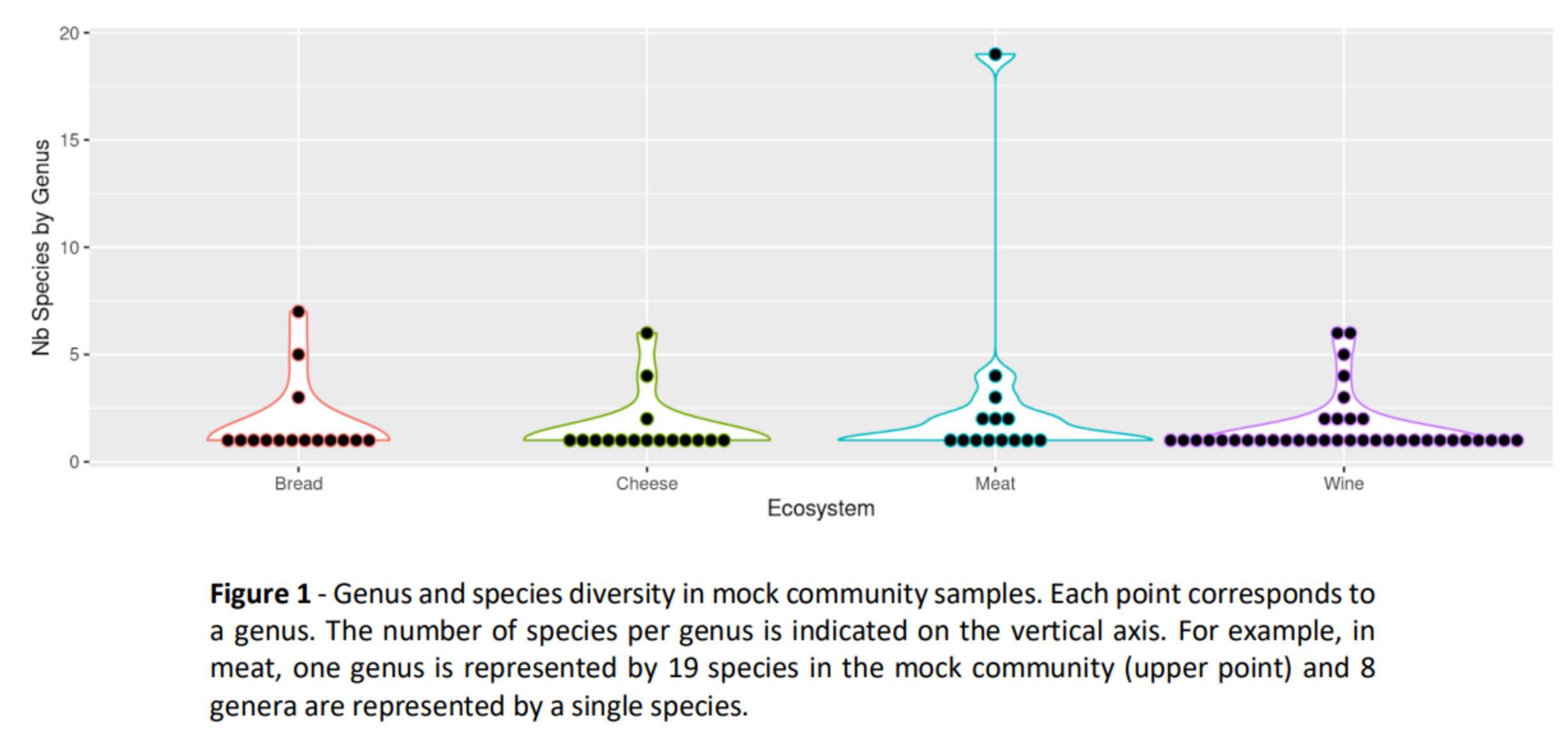
Methods
- For each food environment (bread, wine, cheese, fermented meat), two different mock communities were prepared, a “DNA” mock community and a “PCR” mock community.
- All mock community samples were prepared in triplicate.
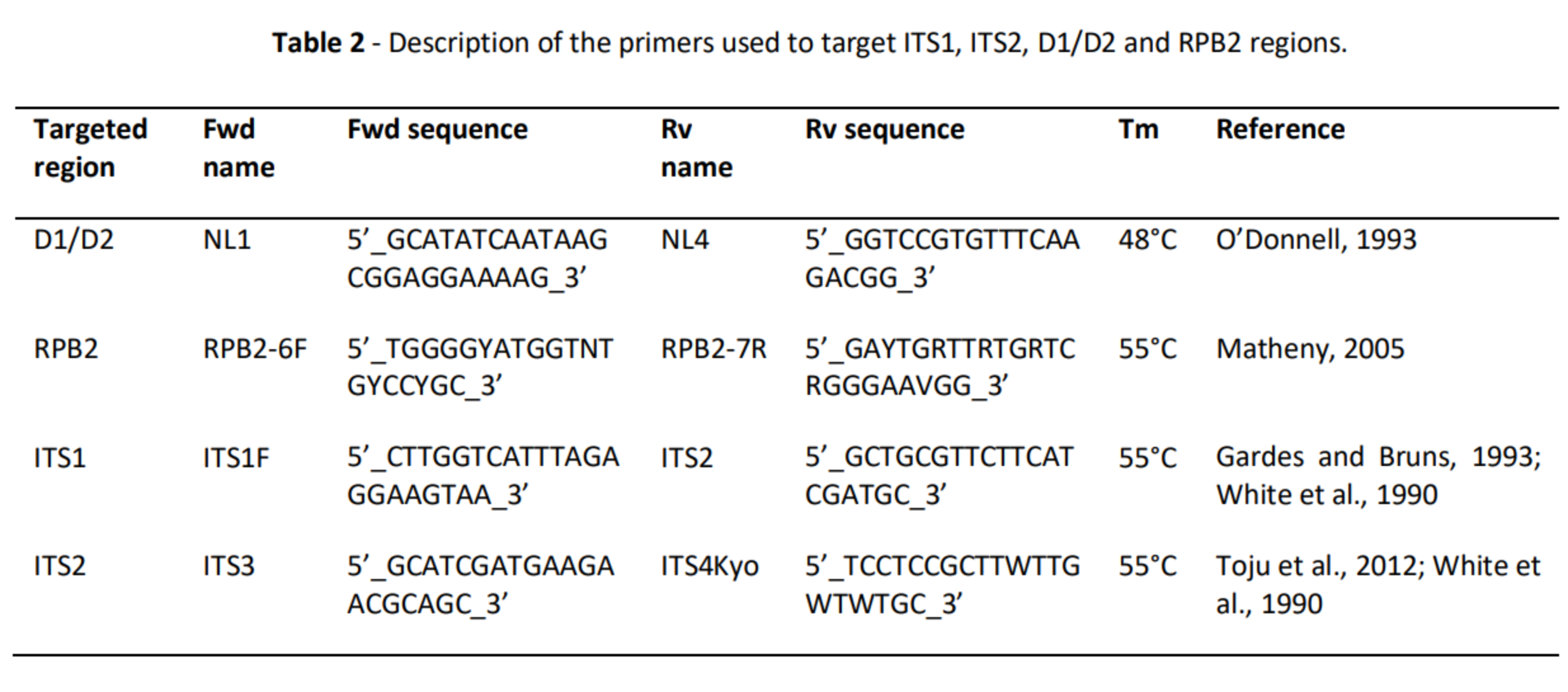

Methods
- Each of the 469 sequences used in mock communities was identified (from INRAE labs, public genomes, dedicated databanks…) (Claire Vincent, M2 internship 2020, MaIAGE) and is part of the reference databank used for bioinformatics
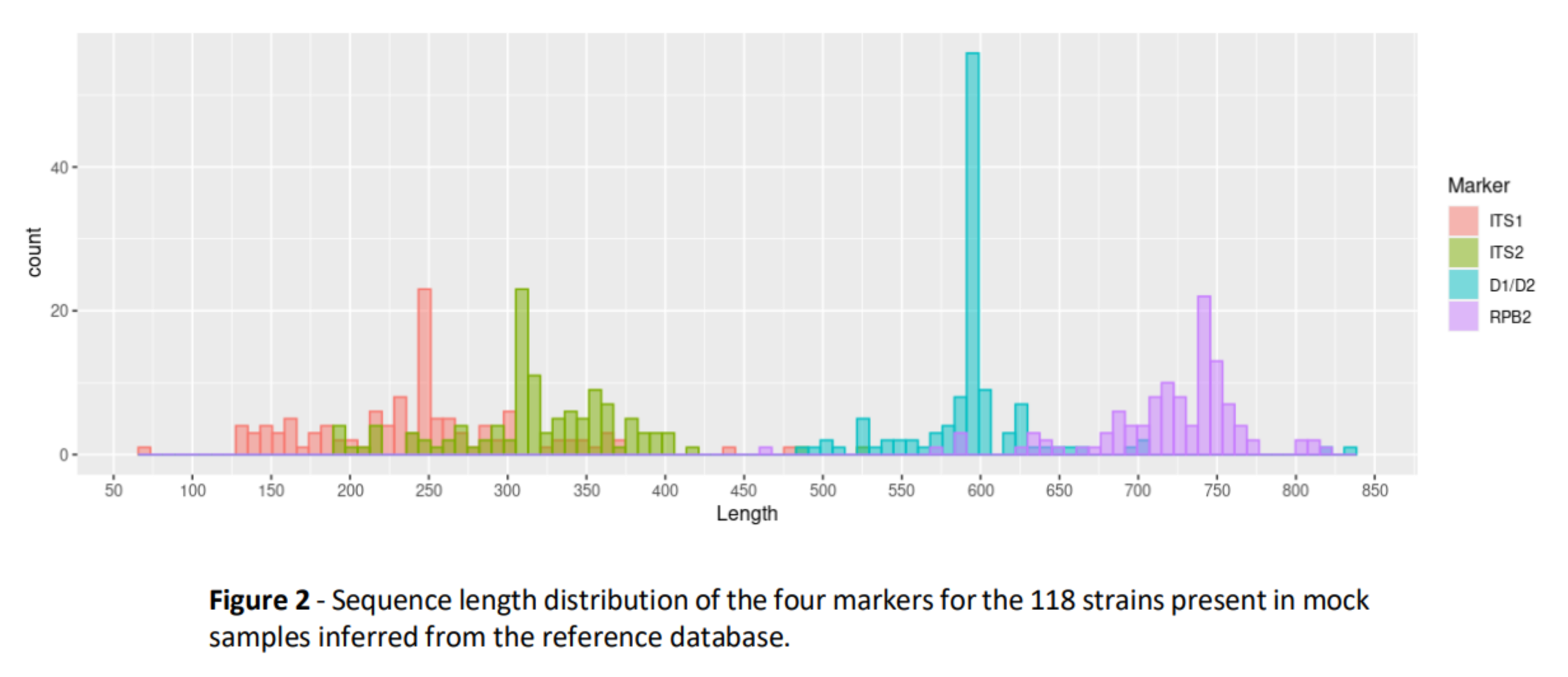
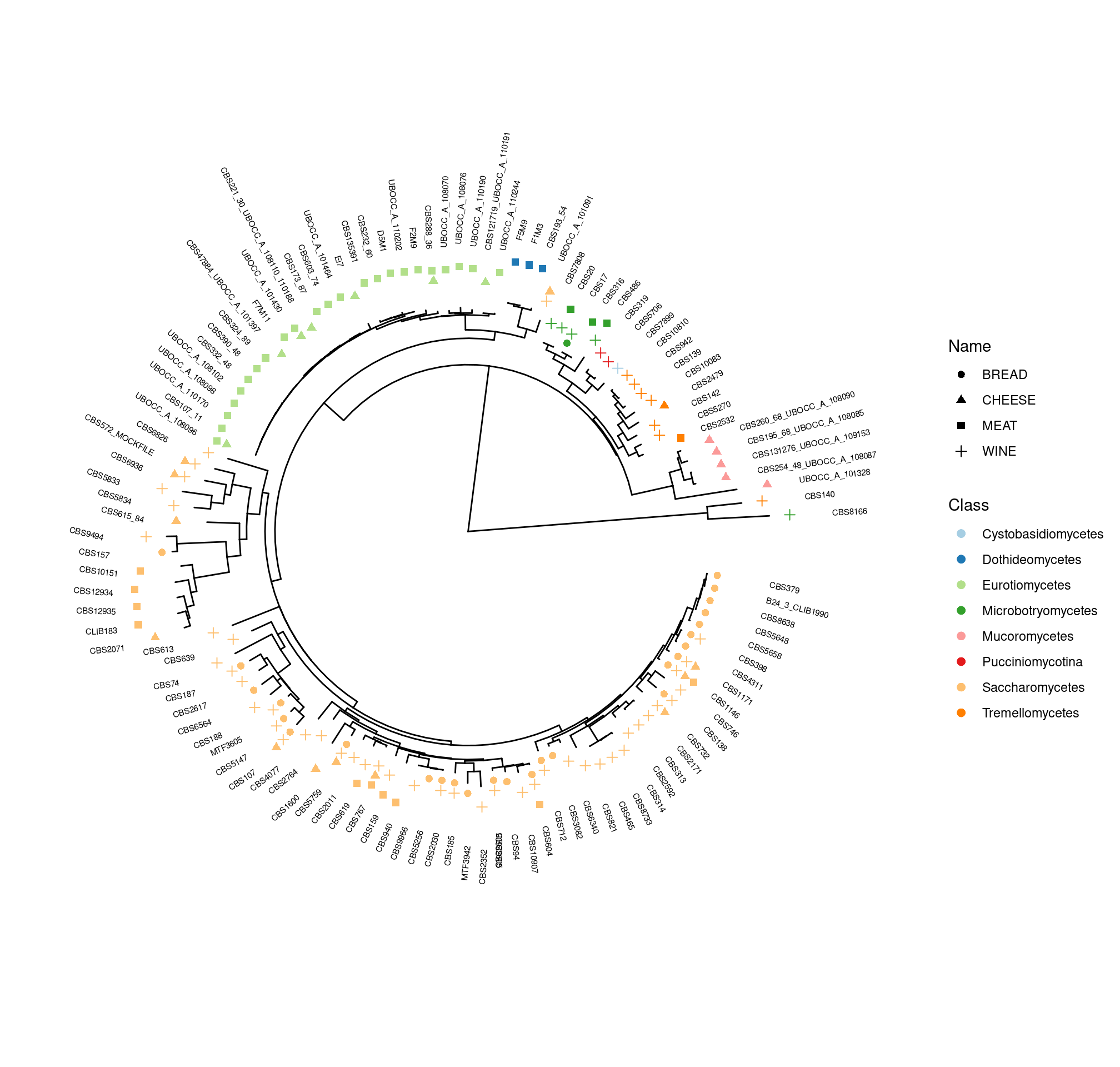
Benchmark
- Bioinformatics tools
- FROGS
- USEARCH
- DADA2 se / pe
- QIIME2 se / pe
- DADA2_FROGS (mix both tools)
- Metrics
- Divergence rate (Bray-curtis distance between observed and expected)
- FN (taxa not recovered)
- FP (supplementary taxa)
- TP (expected and recovered taxa)
- Precision (TP/(TP+FP)) and Recall rate (TP/(TP+FN))
- Perfectly identified sequences (strictly identical sequence to reference)
Benchmark
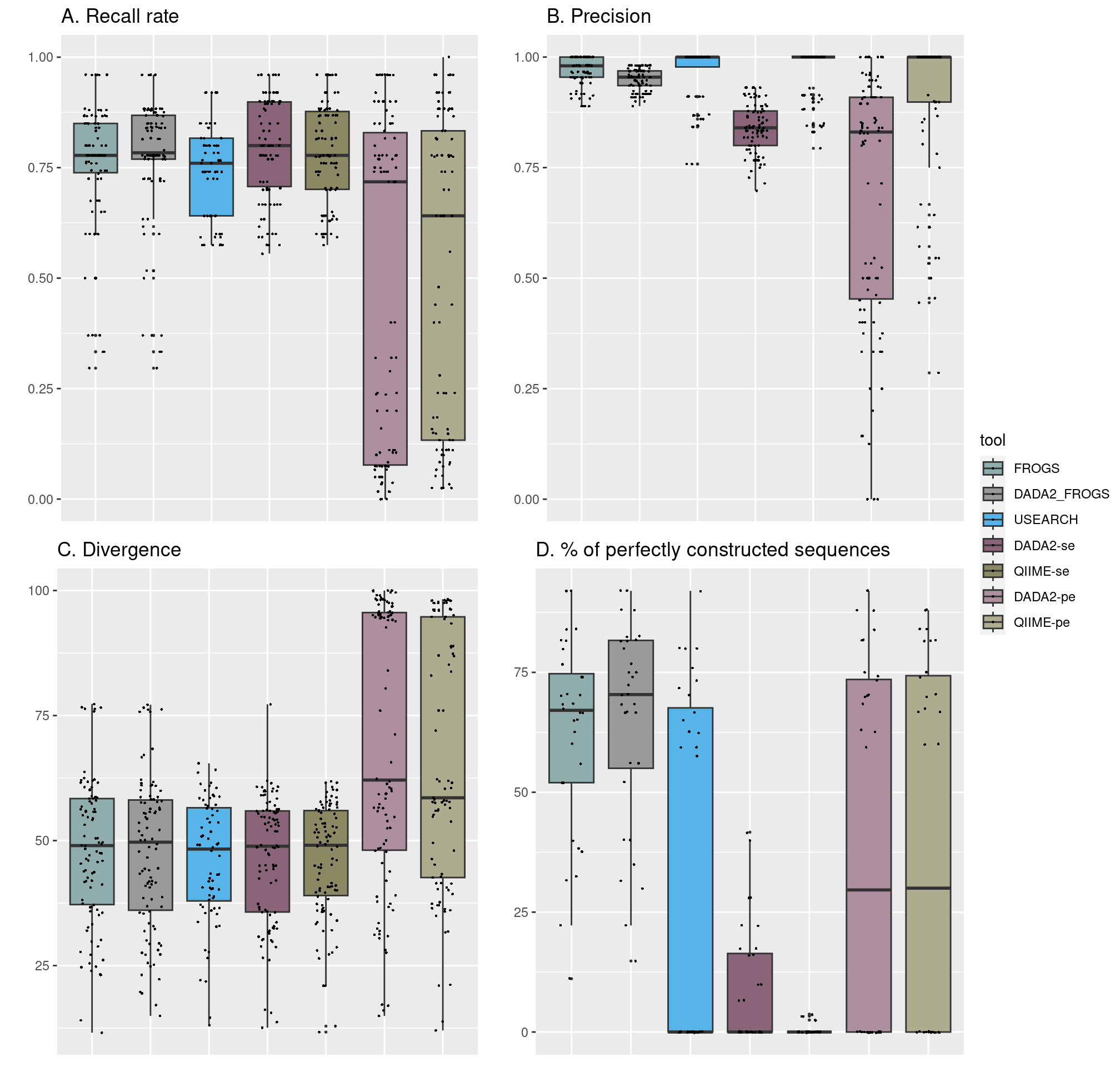
- A. Recall rate lower for pe tools
- B. DADA2 less precise than others
- C. Higher divergence for pe tools
- D. FROGS and DADA2_FROGS better
Benchmark
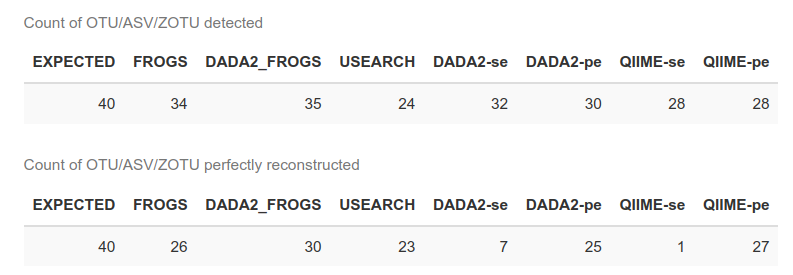
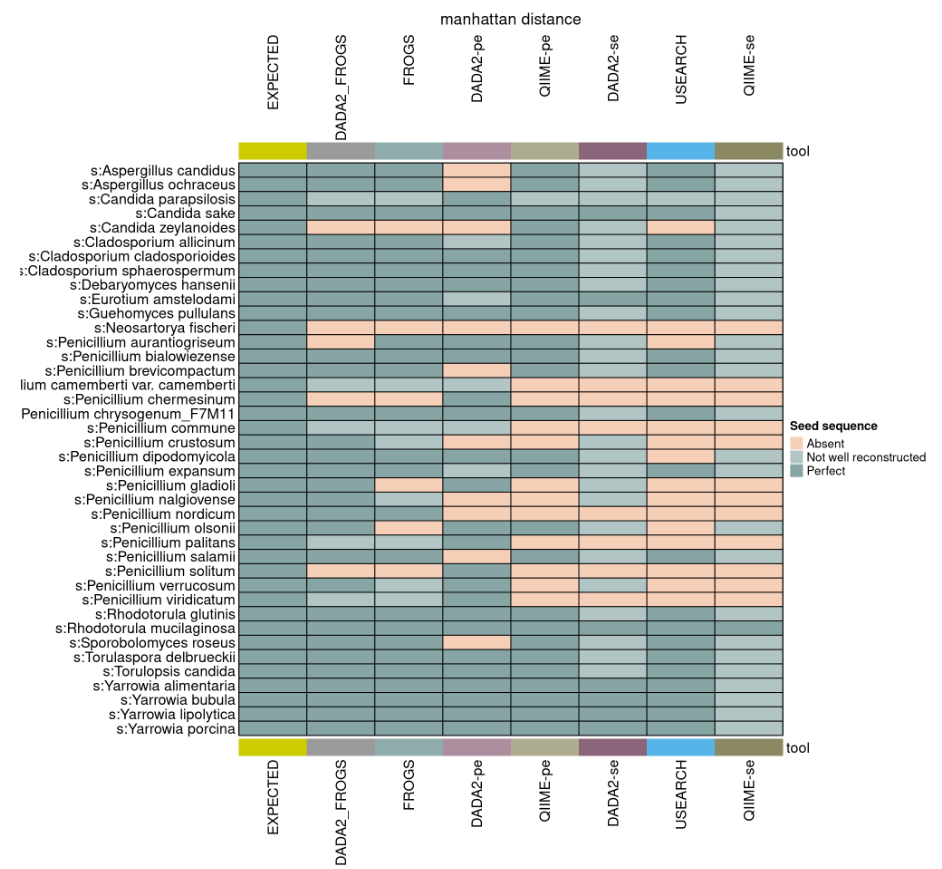
Conclusion 1
DADA2_FROGS is the best approach on mock communities
Marker choice on mock communities
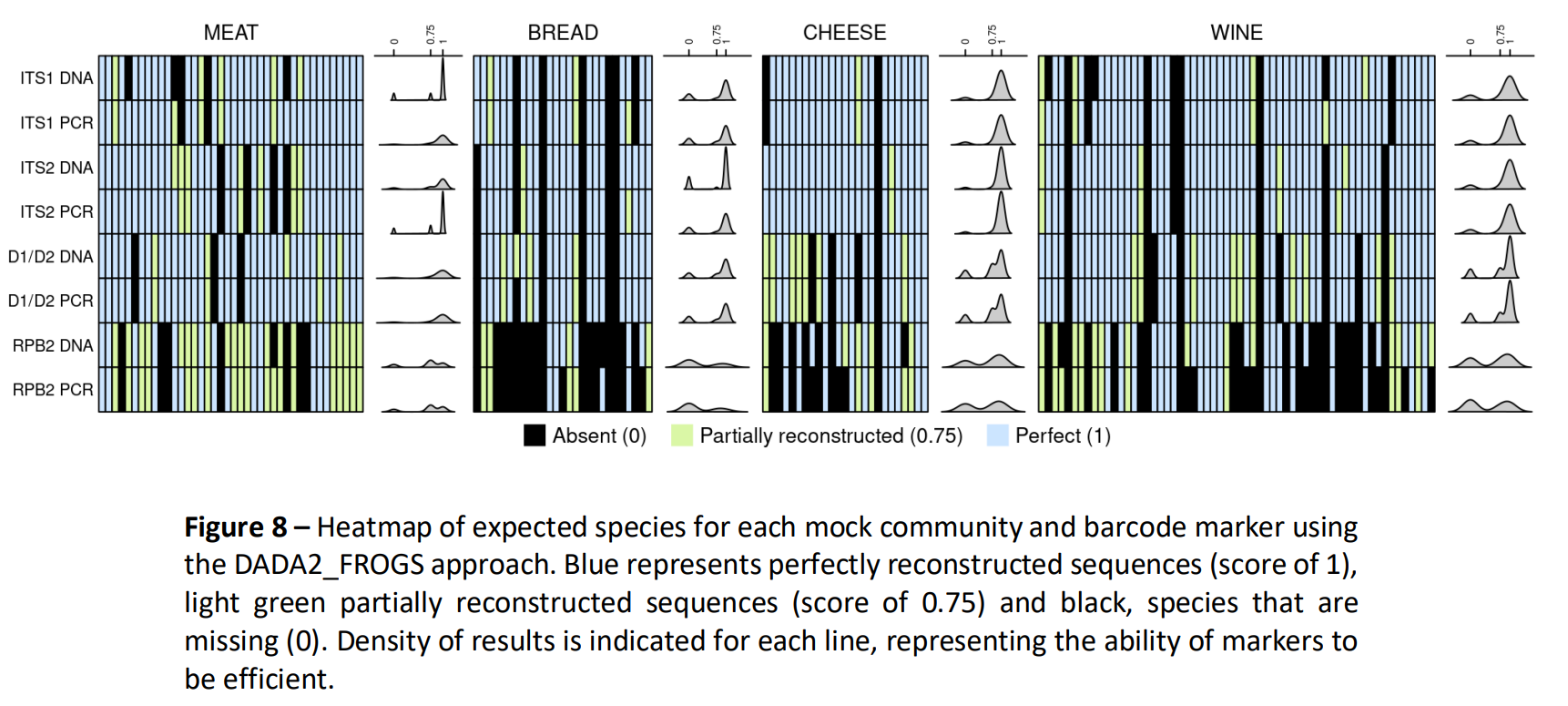
Conclusion 2
RPB2 not suitable based on mock communities
Real samples analysis
- Real samples analyzed with the DADA2_FROGS approach
- Reference databanks consituted of the most used databank + our references
- UNITE 8.2 for ITS; Silva v138 for D1/D2 and NCBI nt for RPB2
- Manual curation/expertise of ASVs obtained
- 2,359 ASVs manually curated and available to the community
Importance of the reference databanks
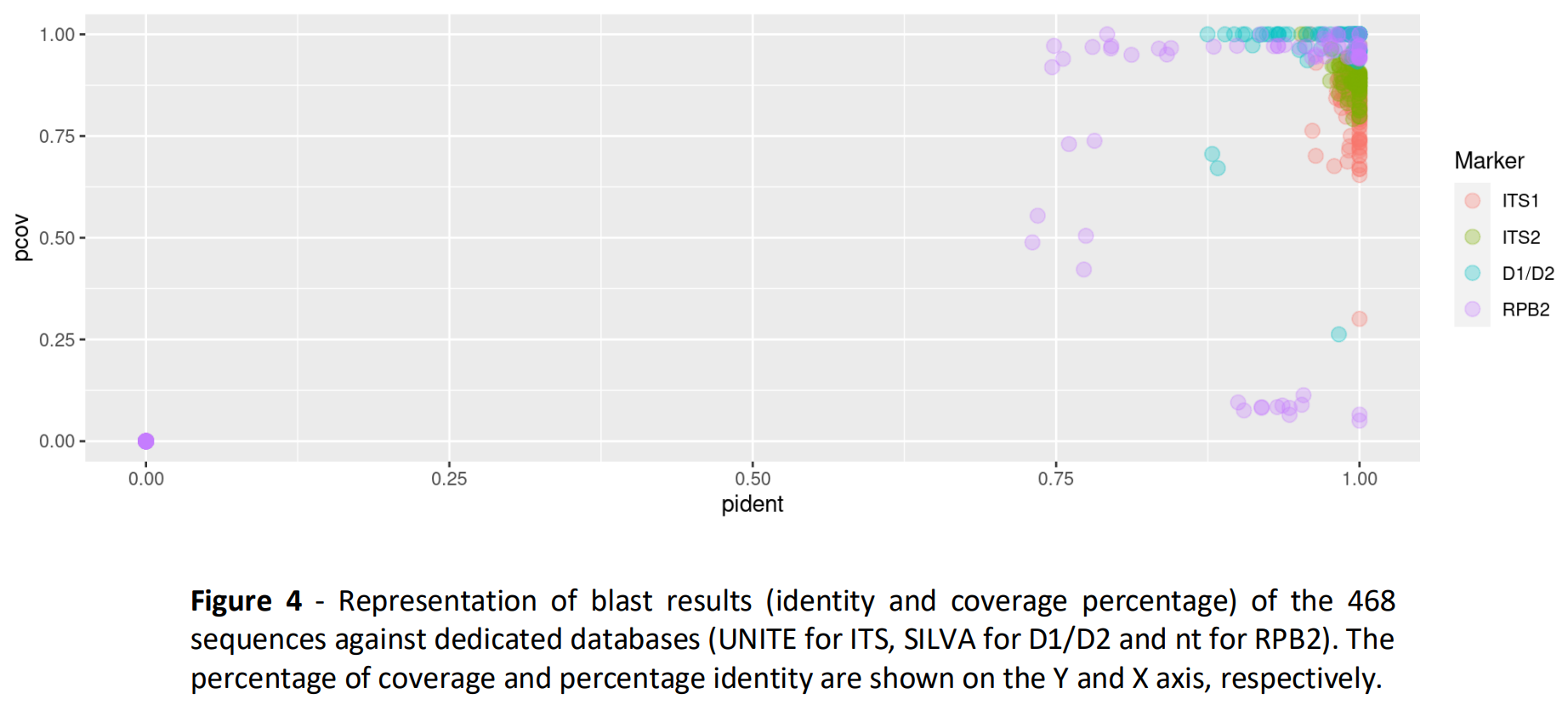
Real samples analysis
Choice of the best marker
- Are key species well identified?
- Is there contamination?
- i.e. amplification of plant DNA for D1/D2. Automatically removed with ITS1/ITS2 data with ITSx tool which is part of FROGS
| Bread | Cheese | Meat | Wine | |
|---|---|---|---|---|
| Best marker | ITS1 | ITS2 | ITS2 | ITS2 |
Conclusion 3
ITS markers performed better than D1/D2, as they are better represented in public databases and have better specificity to distinguish species
Conclusion 4
No generic recommendation for all fermented food types can be made
Conclusions
A long-term project
- COVID-19, other priorities…
- Reclaiming leadership to mobilize the troops
- Setting up collaborative tools
- Code deposited on forgeMIA
- continuous deployment of analyses reports
- reproducible analyses, a lot of time saving!
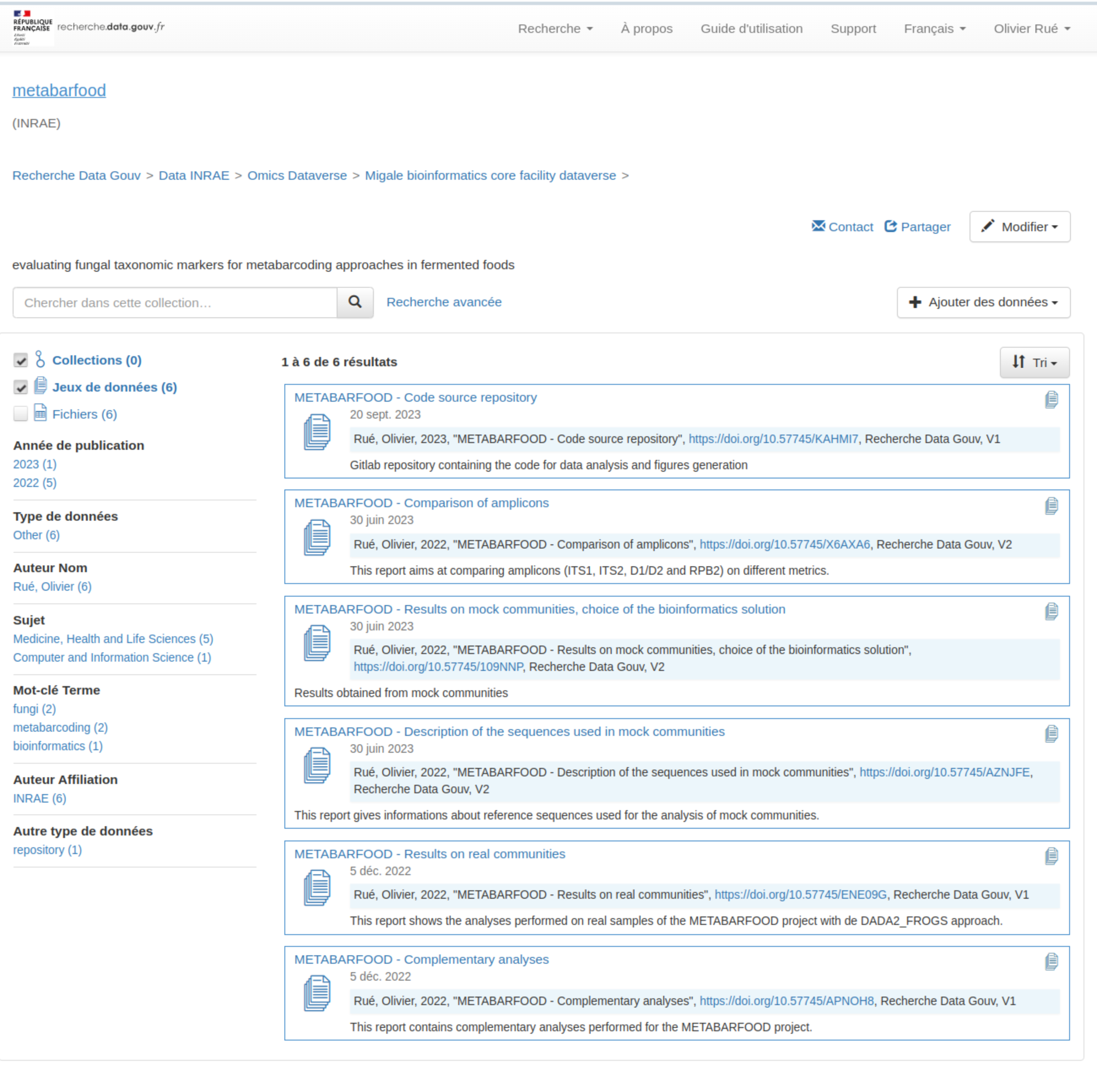
- Dedicated dataverse on Recherche Data Gouv / Code archived on HAL ↔︎ Software Heritage
- DADA2_FROGS approach being integrated in FROGS v.5.0
- Publication in Peer Community Journal in 2023

Thanks for your attention
References
1. Escudié F, Auer L, Bernard M, Mariadassou M, Cauquil L, Vidal K, et al. FROGS: Find, Rapidly, OTUs with Galaxy Solution. Bioinformatics. 2018;34:1287–94. doi:10.1093/bioinformatics/btx791.
2. Bernard M, Rué O, Mariadassou M, Pascal G. FROGS: a powerful tool to analyse the diversity of fungi with special management of internal transcribed spacers. Briefings in Bioinformatics. 2021;22. doi:10.1093/bib/bbab318.

Séminaire interne MaIAGE