Analyse de données métagénomiques 16S
Module 20
June 8, 2026
Introduction
Practical informations
- 9h00 - 17h00
- 2 breaks morning and afternoon
- Lunch at INRAE restaurant (not mandatory)
- Questions are strongly encouraged
- Everyone has something to learn from each other
Better know you
Who are you?
- Institution / Laboratory / position
What is your scientific topic?
- Studied ecosystem
- Scientific question
- Experimental design
What is your background?
- Already treated shotgun data?
- Background in bioinformatics?
- Background in biostatistics?
Better know us
Data analysis service
https://documents.migale.inrae.fr/data-analysis.html
- We are specialized in genomics/metagenomics
- 5 Bioinformaticians and 2 Statisticians
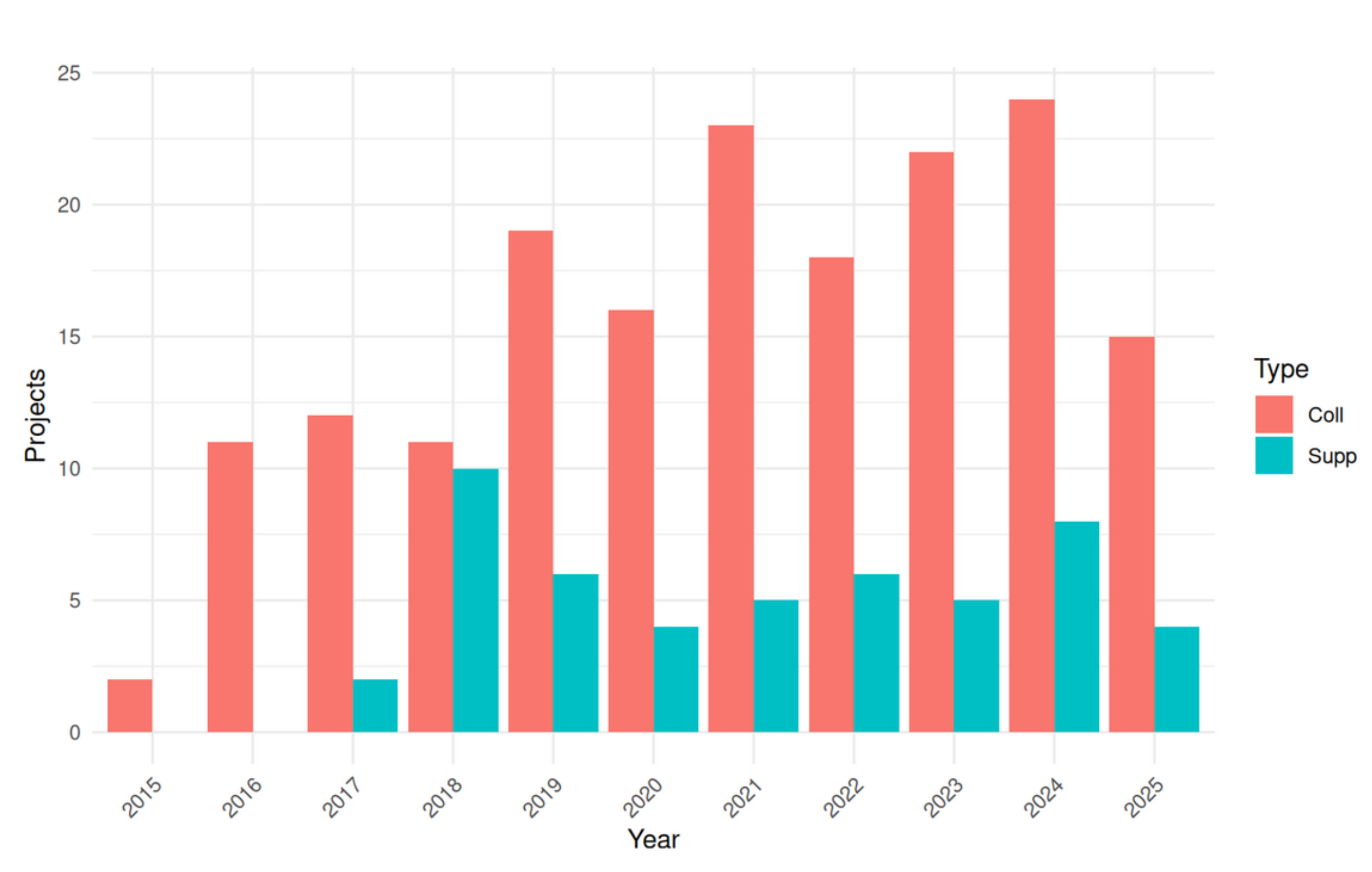
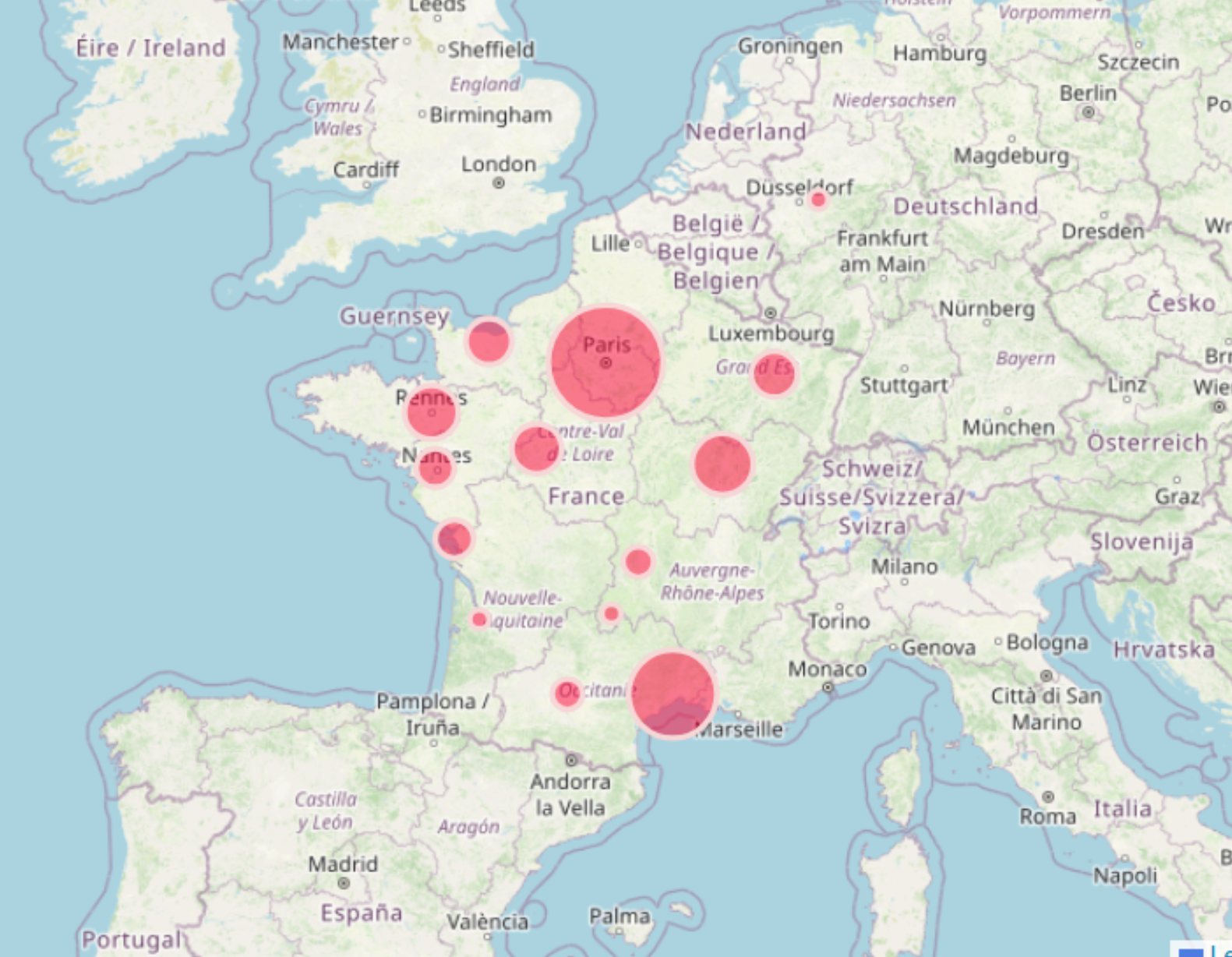
- More than 160 projects since 2016
- LRQA certified process
- 2 types of services
- Classical collaboration (we perfom the analyses)
- Accompaniment (we help you do the analysis yourself)
Aim of this training
After this 4 days training, you will:
- Know the outlines, advantages and limits of amplicon sequencing data analysis
- Be able to use FROGS (through Galaxy) and phyloseq (through easy16S) tools on the training data set
- Be able to identify tools and parameters adapted to your own analyses
Aim of this training
Program
DAY 1
- Introduction
- Introduction to amplicon analysis
- Introduction to Galaxy
- Quality control
- FROGS (1)
DAY 2
- FROGS (2)
- FROGSfunc
DAY 3
- Introduction
- Easy16S
- Composition
- \(\alpha\) and \(\beta\) diversities
- Ordination
DAY 4
- PERMANOVA and hypothesis tests
- Differential abundance
- Train on your own dataset or on another provided dataset
Program
Training with Easy16S
DAY 3
- Introduction
- Easy16S
- Composition
- \(\alpha\) and \(\beta\) diversities
- Ordination
DAY 4
- PERMANOVA and hypothesis tests
- Differential abundance
- Train on your own dataset or on another provided dataset
Microbiome tools
Aims (1/2)
Become familiar with {phyloseq} [2] R package and {Easy16S} [3] Shiny Web Application for the analysis of microbiome datasets.
Exploratory Data Analysis
\(\alpha\)-diversity: how diverse is my community?
\(\beta\)-diversity: how different are two communities?
Use a distance matrix to study structures:
- Hierarchical clustering: how do the communities cluster?
- Permutational ANOVA: Communities structured by some environmental factor?
Aims (2/2)
Become familiar with {phyloseq} [2] R package and {Easy16S} [3] Shiny Web Application for the analysis of microbiome datasets.
Visual assessment of the data
- bar plots: what is the composition of each community?
- Multidimensional Scaling: how are communities related?
- Heatmaps: are there interactions between species and (groups of) communities?
- Differential Abundances: which taxa are differentially abundant?
phyloseq and companion tools
phyloseq R package [2] from Bioconductor
- importing data from a variety of common formats
- preprocessing data (storing, filtering, subsetting, transforming…)
- performing analysis and graphics to explore microbiome profils
Companion tools
- Customs functions designed to enhance core functionality
{phyloseq-extended}[4] - Community ecology functions:
{vegan},{ade4},{picante} - Tree manipulation:
{ape} - Orchestrating Microbiome Analysis with Bioconductor
{mia}
easy16S (1/2)
Shiny Web Application [3]
Strengths
- specifically designed for biologists, data analysts and traineers
- facilitates exploratory microbiome data analysis, data visualization, and statistical analysis
- based on the
{phyloseq}data structure [2] - data visualization with
{ggplot2}
easy16S (2/2)
Shiny Web Application [3]
Content
- Key tables constituting the phyloseq object;
- Metadata visualization using esquisse (Meyer & Perrier, 2018);
- Taxonomic composition barplot;
- Rarefaction curves;
- Abundance heatmap;
- Richness within a sample (α-diversity): table, scatterplot and ANOVA;
- Dissimilarity between samples (β-diversity): table, sample heatmap, sample clustering,
- MultiDimensional Scaling and Multivariate ANOVA;
- Principal Component Analysis;
- Differential abundance analysis.
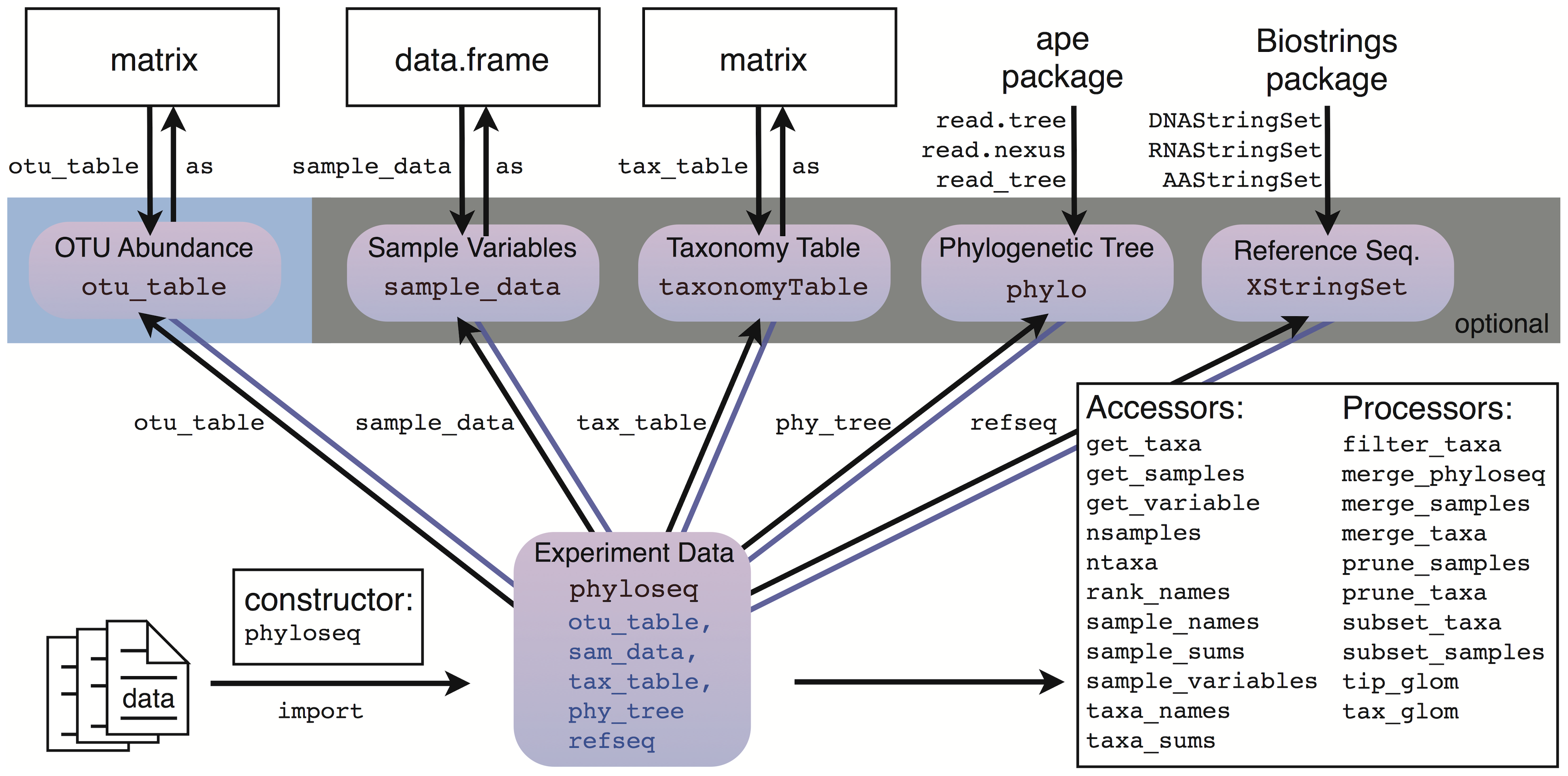
Phyloseq data structure (1/)
Phyloseq data structure (2/)
OTU/ASV abundances
Phyloseq data structure (3/)
sample variables : reflectexperimental desing
Phyloseq data structure (4/)
taxonomy table
Phyloseq data structure (5/)
Phylogenetic Tree
Reference Seq
Description microbiome dataset
practice
References


Module 20 - Metabarcoding