k__Bacteria;p__Actinobacteria;c__Coriobacteriia;o__Coriobacteriales;f__Coriobacteriaceae;g__Slackia;s__Slackia_heliotrinireducenCreate your own FROGS databank
Introduction
FROGS
It allows to analyze large sets of DNA amplicons sequences accurately and rapidly, essential for microbe community studies.
Among the many tools available, a tool called taxonomic_affiliation returns taxonomic affiliation for each ASV using Blast
In this document, we show you how to make your own databank for FROGS. You will need to have rdp_classifier-2.13
How to use a custom databank?
Sometimes, you need to use a databank which is not availabe among those we provide. Depending on whether you use FROGS from the command line or via Galaxy, the problem is not the same.
Command line
In command line, you need to choose the affiliation databank to use, by indicating the sequence reference fasta file with the option --reference. You can therefore use your own databank, on condition that it is well formatted!
Galaxy
With Galaxy, you have to chose a databank among a list of available databanks. You can’t chose another one without asking administrators to add it.
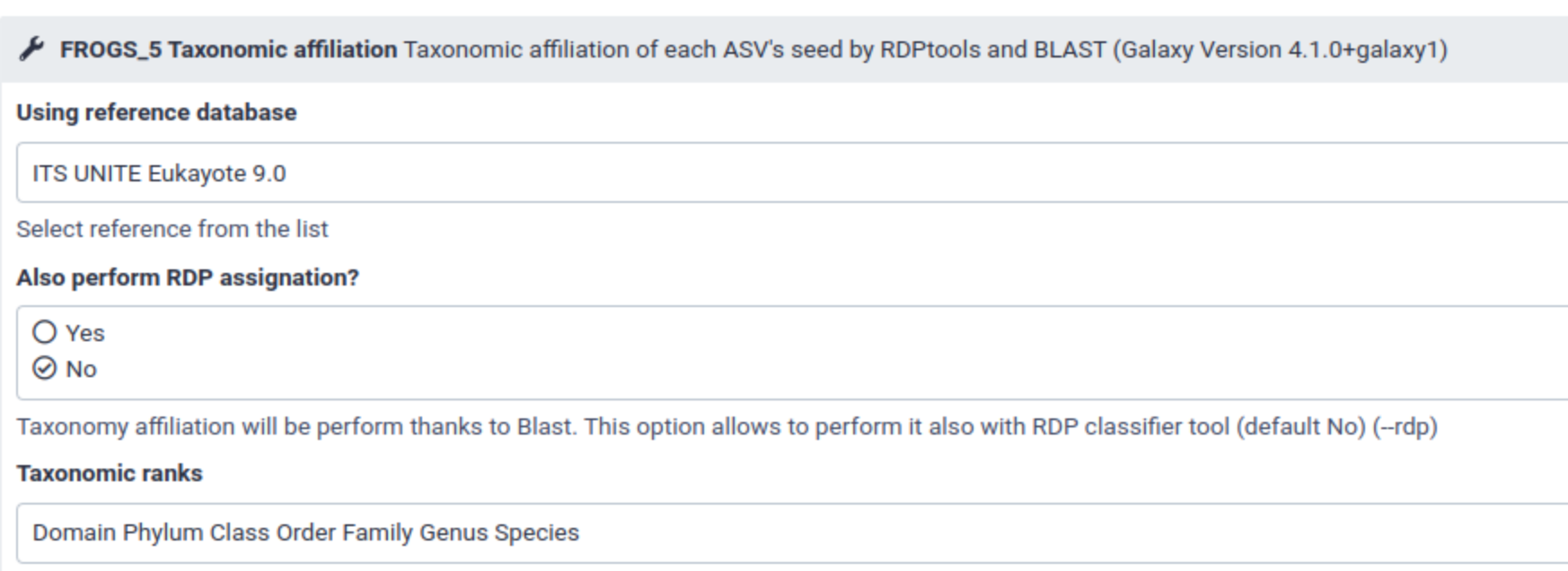
A databank available in Galaxy is accessible by all the users of this instance.
Prerequites for a FROGS databank
Four informations are mandatory to build a FROGS databank:
- a name and a version
- some nucleic sequences (the targeted region of interest)
- a unique identifier for each sequence
- a taxonomy for each sequence
Name and version
As the content of the banks is constantly evolving, it is essential to specify the version. It can be a number, a date…
The sequence
Only A, C, G, T or N characters are authorized (nucleotidic sequence). The length is not important.
The header of the sequences
Each sequence must have a unique identifier.
The taxonomic ranks
A FROGS databank needs each sequence to have the exact same number of taxonomic ranks. The classical 7 ranks are:
| Phylum | Class | Order | Family | Genus | Species |
but can also be:
| Phylum | Class | Order | Family | Genus | Species | Strain |
The taxonomy can be written following the Greengenes taxonomy:
or not:
Bacteria;Actinobacteria;Coriobacteriia;Coriobacteriales;Coriobacteriaceae;Slackia;Slackia_heliotrinireducenPrepare the raw files
To format a databank, you will need to create two files. One containing the taxonomic ranks (called here taxonomies.tsv), the other one containing the sequences (sequences.fasta).
Example:
taxonomies.tsv
SEQ1 Bacteria;Actinobacteria;Coriobacteriia;Coriobacteriales;Coriobacteriaceae;Slackia;Slackia_heliotrinireducen
SEQ2 Bacteria;Actinobacteria;Coriobacteriia;Coriobacteriales;Coriobacteriaceae;Slackia;Slackia_exiguasequences.fasta
>SEQ1
ATTCCTTGGAGGTGAGGTGCGGGCGCGAG
>SEQ2
TTTCCTTGGAGGTGAGGTGCGGGCGCCCCYou can add the databank name and version in the filename.
DB_NAME="MYDB"
DB_VERSION="MYVERSION"
mkdir ${DB_NAME}_${DB_VERSION}
FASTA=${DB_NAME}_${DB_VERSION}.fasta
TAX=${DB_NAME}_${DB_VERSION}.taxIndex files for FROGS
Check the consistency of the raw files
First, a verification is necessary to ensure that the files are as expected.
grep -c '>' $FASTA
2
wc -l $TAX
2Check all sequences have a taxonomy
grep '>' $FASTA | sed 's/>//' | awk -v T=$TAX 'BEGIN{while(getline<T>0){tab[$1]=1}}{if(tab[$1]!=1){print $1}}' > no_tax.txt
wc -l no_tax.txt
0Check all taxonomies harbor the same ranks number
cut -f 2 $TAX |awk -F ";" '{print NF}' | sort -u
7The result is 7 if your taxonomies have 7 ranks, but can be 6, 8…
Check the N rate
cat $FASTA |awk -F "N" '{if(substr($1,0,1) == ">"){split($1,tab," "); id=tab[1]}else{count=NF-1; print id"\t"count"\t"length($0)"\t"count*100/length($0)}}' > count_N.txt
cat count_N.txt | awk '{c++; if($4>40){m++}}END{print m"/"c"="m*100/c"%"}'It is a recommandation of Blast to have a small proportion of N’s in the sequences.
RDP files
The script fasta2RDP.py (available in FROGS sources) allows to generate mandatory files for RDP.
fasta2RDP.py -d $FASTA -t $TAX -r R1 R2 R3 R4 R5 R6 R7 --rdp-taxonomy ${DB_NAME}_${DB_VERSION}/${DB_NAME}_${DB_VERSION}.tax --rdp-fasta ${DB_NAME}_${DB_VERSION}/${DB_NAME}_${DB_VERSION}.fastaThe -r option allows to precise the rank names. These names do not have any impact on the rank names used in FROGS. However, there must be as many as the number of ranks.
conda activate frogs-4.1.0
conda_lib_dir=`echo $(dirname $(dirname $(which preprocess.py)))/libexec`
export PATH=$conda_lib_dir:$PATH
#export PATH="/home/orue/work/GIT/FROGS2023OK/libexec":$PATH
export PYTHONPATH=`echo $(dirname $(dirname $(which preprocess.py)))/lib`:$PYTHONPATH
fasta2RDP.py -d $FASTA -t $TAX -r R1 R2 R3 R4 R5 R6 R7 --rdp-taxonomy ${DB_NAME}_${DB_VERSION}/${DB_NAME}_${DB_VERSION}.tax --rdp-fasta ${DB_NAME}_${DB_VERSION}/${DB_NAME}_${DB_VERSION}.fasta
conda deactivateThe following commands allow to check if problematic taxa are found. Problematic taxa are:
- duplicated taxon
- a taxon name used at different ranks or used at the same rank but with different kinship
cut -f 2 -d '*' ${DB_NAME}_${DB_VERSION}/${DB_NAME}_${DB_VERSION}.tax | cut -f 1 -d ' ' | sort | uniq -c | awk '$1!=1 && $2 !="unclassified"' > problematic_taxon
if [[ -e problematic_taxon_and_rank_number ]]
then
rm problematic_taxon_and_rank_number
fi
for i in `awk '{print $2}' problematic_taxon `
do
exp=`echo \"\*$i \[id\"`
grep "\*"$i" \[id" ${DB_NAME}_${DB_VERSION}/${DB_NAME}_${DB_VERSION}.tax | sed 's/ \[id/*/'| awk -F "*" '{print $2,$5}' >> problematic_taxon_and_rank_number
doneThen, the tool classifier.jar of the rdp tools is run:
java -Xmx60g -jar classifier.jar train -o . -s ${DB_NAME}_${DB_VERSION}/${DB_NAME}_${DB_VERSION}.fasta -t ${DB_NAME}_${DB_VERSION}/${DB_NAME}_${DB_VERSION}.taxconda activate rdp_classifier-2.13 && java -Xmx60g -jar /usr/local/genome/Anaconda3/envs/rdp_classifier-2.13/share/rdp_classifier-2.13-1/classifier.jar train -o . -s ${DB_NAME}_${DB_VERSION}/${DB_NAME}_${DB_VERSION}.fasta -t ${DB_NAME}_${DB_VERSION}/${DB_NAME}_${DB_VERSION}.tax && conda deactivateThe following files must be created in the ${DB_NAME}_${DB_VERSION}/ directory:
bergeyTrainingTree.xml
logWordPrior.txt
wordConditionalProbIndexArr.txt
genus_wordConditionalProbList.txtA file called ${DB_NAME}_${DB_VERSION}/${DB_NAME}_${DB_VERSION}.fasta.properties needs to be added manually. You can copy its content (with the correct version of rdp):
# Sample ResourceBundle properties file
bergeyTree=bergeyTrainingTree.xml
probabilityList=genus_wordConditionalProbList.txt
probabilityIndex=wordConditionalProbIndexArr.txt
wordPrior=logWordPrior.txt
classifierVersion=RDP Naive Bayesian rRNA Classifier Version 2.5, May 2012 BLAST files
For BLAST, the tool makeblastdb is used:
makeblastdb -in ${DB_NAME}_${DB_VERSION}/${DB_NAME}_${DB_VERSION}.fasta -dbtype nuclconda activate blast-2.13.0 && makeblastdb -in ${DB_NAME}_${DB_VERSION}/${DB_NAME}_${DB_VERSION}.fasta -dbtype nucl && conda deactivateTest your databank
You can download test files from FROGS github repository:
wget https://github.com/geraldinepascal/FROGS/raw/dev/tools/taxonomic_affiliation/data/swarm.fasta
wget https://github.com/geraldinepascal/FROGS/raw/dev/tools/taxonomic_affiliation/data/swarm.biomand run taxonomic_affiliation.py:
conda activate frogs-4.1.0 && taxonomic_affiliation.py --rdp --reference ${DB_NAME}_${DB_VERSION}/${DB_NAME}_${DB_VERSION}.fasta --input-biom swarm.biom --input-fasta swarm.fasta --output-biom affiliation.biom --summary affiliation.summary && conda deactivateExample files come from a 16S analysis. You may not have affiliations in the output files if your marker gene of interest is not 16S. Nevertheless, the program should not return an error
Availiablily of your databank
Metadata
It is important to add a licence and a README file to explain how was built your databank and how it can be reused by others.
touch ${DB_NAME}_${DB_VERSION}/LICENCE.txt
# and write informations
touch ${DB_NAME}_${DB_VERSION}/readme.txt
# and write informationsFinally, create an archive containing all files:
tar -cvzf ${DB_NAME}_${DB_VERSION}.tar.gz ${DB_NAME}_${DB_VERSION}Display it in LEAP
If you want to make your databank available in LEAP, you can ask us. It will be done several days after availability in FROGS.
